G2C::PhosphoProteomics
Neurotransmitters Drive Combinatorial Multistate Postsynaptic Density Networks
Coba MP1, Pocklington AJ2, Collins MO3, Kopanitsa MV1, Uren RT1, Swamy S3, Croning MD1, Choudhary JS3, Grant SG1.
Author email: sg3@sanger.ac.uk
- Genes to Cognition, Wellcome Trust Sanger Institute, Hinxton, Cambridgeshire CB10 1SA, UK.
- Institute for Adaptive and Neural Computation, Division of Informatics, University of Edinburgh, 5 Forrest Hill, Edinburgh EH1 2QL, UK.
- Proteomic Mass Spectrometry, Wellcome Trust Sanger Institute, Hinxton, Cambridgeshire CB10 1SA, UK.
Synapse phosphoproteome mapping resource
The synapse is a subcellular compartment housing a remarkably diverse set of signaling proteins. Neurotransmitters activate phosphorylation and dephosphorylation of intracellular proteins and these processes are at the centre of understanding the regulation of synaptic transmission, plasticity, forms of behaviour including learning and addiction, as well as contributing to numerous diseases.
We have generated a resource for mapping the complexity of postsynaptic signaling by combining mass spectrometry data from mouse brain, which identifies the state of phosphorylation of specific sites on substrates, with peptide array mapping, which identifies the kinases responsible for phosphorylating specific sites. Details of the data and methods are found in Coba et al, (Mapping postsynaptic phosphoproteome networks: integration and switching mechanisms underlying NMDA receptor signaling, submitted). This resource is embedded in G2Cdb so that data can be linked to phenotype data in mice and humans and other datasets in G2Cdb (and its external links). A set of frequently asked questions (FAQs) is presented to illustrate some of the major uses of the resource, and it is recommended that users click on the FAQs and then bookmark specific tables.
Data resources
1. Is this substrate phosphorylated in vivo, where are the sites, which kinases phosphorylate that site?
Unable to read file: psd-95.pngGo to Table 1 and scroll down for the name or ID of the protein (example screen below). For each protein substrate all the sites that were mapped, the number of kinases and their specific names for each site are shown. Also included is literature reference (PubMed PMID or link) and position of site as referred to in this publication.
Using PSD-95 as an example substrate, there are 9 reported phosphorylation sites (T19, S25, S35, T287, T290, S295, S415, S418, Y432).
2. What are the synaptic substrates for this kinase?
Unable to read file: camkii.pngGo to Table 2 to see list of kinases and the substrates and sites. For example, CamkII (which is important in synaptic plasticity and learning) has 16 identified substrates.
3. Which proteins are phosphorylated and dephosphorylated when the NMDA receptor is stimulated?
Go to Table 3 to see list of substrates and if they were found to be dephosphorylated or phosphorylated after NMDA stimulation of hippocampus.
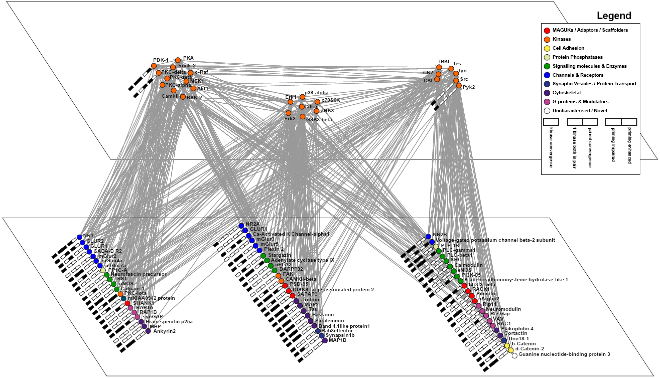
4. What does a synaptic phosphoproteome network look like?
The kinase-substrate data can be graphed to show the complexity of the postsynaptic phosphorylation networks.
5. How do I find out if this kinase or substrate is required for synaptic plasticity in mutant mice?
G2Cdb is an integrated database for studying neurobiology and contains data on mouse knockouts and synaptic plasticity. This is a curated database of literature and data directly deposited from the Genes to Cognition mutant mouse phenotyping pipeline. Links from the substrates and kinases described in these synapse phosphoproteome pages to this data can be found via the G2Cdb gene name links provided in the tables.
6. Is this kinase or substrate involved in a human disease?
G2Cdb also contains information on synaptic genes and their involvement in disease as well as links to OMIM. We have focussed the disease curated literature on components of the postsynaptic proteome, particularly components of the NRC/MASC complexes. Links from the substrates and kinases described in these synapse phosphoproteome pages to this data can be found via the G2Cdb gene name links provided in the tables.
7. Where do I find a detailed description of the methods and results for mapping synapse phosphorylation networks?
Table 1. Substrate phosphorylation sites and their cognate kinases
The protein substrate name and identifiers (G2Cdb gene ID and Uniprot accession) are shown. The specific site in that substrate is indicated (e.g. S184) and where a sequence is shown (e.g. FYYEILNSPEKACSL) this indicates that the specific phosphoacceptor residue was not unambiguously defined within that sequence. The number of kinases phosphorylated that site/sequence and names is shown. Links to literature and mouse genetic and human disease data is obtained via the G2Cdb gene link.
Table 2. Kinases and their substrates
The table is organised so that the substrates and sites for each kinase is reported. The kinase and protein substrate name and identifiers (G2Cdb gene ID and Uniprot accession) are shown. The specific site in that substrate is indicated (e.g. S184) and where a sequence is shown (e.g. FYYEILNSPEKACSL) this indicates that the specific phosphoacceptor residue was not unambiguously defined within that sequence. Links to literature and mouse genetic and human disease data is obtained via the G2Cdb gene link
| Kinase | G2Cdb gene | Substrate | Substrate UniProt | Site/peptide |
|---|---|---|---|---|
| Akt1 | G00000175 | 14-3-3 zeta | P63101 | S58 |
| Akt1 | G00000175 | Calpain-1 | O35350 | S379 |
| Akt1 | G00000175 | Calpain-1 | O35350 | T380 |
| Akt1 | G00000175 | DISKS large-associated protein 2 | Q8BJ42 | S440 |
| Akt1 | G00000175 | GLUR1 | P23818 | S710 |
| Akt1 | G00000175 | GLUR2 | P23819 | S717 |
| Akt1 | G00000175 | IRS1 | P35569 | S24 |
| Akt1 | G00000175 | IRS1 | P35569 | S265 |
| Akt1 | G00000175 | mGlur4a | P31423 | S859 |
| Akt1 | G00000175 | mGlur7a | Q68ED2 | S862 |
| Akt1 | G00000175 | mKIAA0942 protein | Q8CEA6 | TISRIGSTTNPFLDI |
| Akt1 | G00000175 | NR1 | P35438 | S889 |
| Akt1 | G00000175 | NR1 | P35438 | S890 |
| Akt1 | G00000175 | PKC-iota | Q62074 | S254 |
| Akt1 | G00000175 | PSD-95 | Q62108 | DYPTAMTPTSPRRYS |
| Akt1 | G00000175 | RasGap | P50904 | TVDGKEIYNTIRRKT |
| Akt1 | G00000175 | SHANK1 | Q9WV48 | T635 |
| CamkII | G00000016 | 14-3-3 zeta | P63101 | S58 |
| CamkII | G00000016 | Ankyrin2 | Q01484 | PKKSDSNASFLRAAR |
| CamkII | G00000016 | Ankyrin2 | Q01484 | S31 |
| CamkII | G00000016 | Ankyrin2 | Q01484 | S34 |
| CamkII | G00000016 | Brain specific p25a | Q7TQD2 | S19 |
| CamkII | G00000016 | Calpain-1 | O35350 | T380 |
| CamkII | G00000016 | GLUR1 | P23818 | S849 |
| CamkII | G00000016 | GLUR2 | P23819 | S683 |
| CamkII | G00000016 | IRS1 | P35569 | S24 |
| CamkII | G00000016 | mGlur4a | P31423 | S859 |
| CamkII | G00000016 | mGlur7a | Q68ED2 | S862 |
| CamkII | G00000016 | nNOS | Q9Z0J4 | S374 |
| CamkII | G00000016 | nNOS | Q9Z0J4 | TNRLRSESIAFIEES |
| CamkII | G00000016 | NR2B | Q01097 | S1303 |
| CamkII | G00000016 | PKC-delta | P28867 | T50 |
| CamkII | G00000016 | PP1C-A | P62137 | T320 |
| CamkII | G00000016 | PSD-95 | Q62108 | S290 |
| CamkII | G00000016 | SHANK1 | Q9WV48 | KRLFRHYTVGSYDSF |
| CamkII | G00000016 | synGAP | Q9QUH6 | S751 |
| Cdk5 | G00000167 | 14-3-3 zeta | P63101 | S184 |
| Cdk5 | G00000167 | Ankyrin2 | Q01484 | S2439 |
| Cdk5 | G00000167 | Bassoon | O88737 | S1418 |
| Cdk5 | G00000167 | Bassoon | O88737 | S3217 |
| Cdk5 | G00000167 | Ca-Activated K Channel-alpha1 | Q08460 | S711 |
| Cdk5 | G00000167 | CAMKII-beta | P28652 | T321 |
| Cdk5 | G00000167 | cPLA2 | P47713 | S437 |
| Cdk5 | G00000167 | DARPP32 | Q60829 | T34 |
| Cdk5 | G00000167 | DARPP32 | Q60829 | T75 |
| Cdk5 | G00000167 | Drebrin | Q9QXS6 | S141 |
| Cdk5 | G00000167 | FAK | P34152 | S770 |
| Cdk5 | G00000167 | GLUR2 | P23819 | NPSSSQNSQNFATYK |
| Cdk5 | G00000167 | GLUR2 | P23819 | S880 |
| Cdk5 | G00000167 | IRS1 | P35569 | NLHTDDGYMPMSPGV |
| Cdk5 | G00000167 | IRS1 | P35569 | S24 |
| Cdk5 | G00000167 | MAP1B | P14873 | S1775 |
| Cdk5 | G00000167 | mGlur5 | P31424 | S1014 |
| Cdk5 | G00000167 | mGlur5 | P31424 | S1016 |
| Cdk5 | G00000167 | NR1 | P35438 | S889 |
| Cdk5 | G00000167 | NR2A | P35436 | S1232 |
| Cdk5 | G00000167 | Paralemmin | Q9Z0P4 | S157 |
| Cdk5 | G00000167 | Paralemmin | Q9Z0P4 | T141 |
| Cdk5 | G00000167 | Paralemmin | Q9Z0P4 | T145 |
| Cdk5 | G00000167 | PDK-1 | Q9Z2A0 | S384 |
| Cdk5 | G00000167 | PKC-delta | P28867 | EILQGLKYSFSVDWW |
| Cdk5 | G00000167 | PSD-95 | Q62108 | DYPTAMTPTSPRRYS |
| Cdk5 | G00000167 | PSD-95 | Q62108 | S25 |
| Cdk5 | G00000167 | PSD-95 | Q62108 | S290 |
| Cdk5 | G00000167 | PSD-95 | Q62108 | T19 |
| Cdk5 | G00000167 | Rab3effector | Q9JIR4 | S895 |
| Cdk5 | G00000167 | RasGap | P50904 | TVDGKEIYNTIRRKT |
| Cdk5 | G00000167 | SAPAP3 | Q6PFD5 | S58 |
| Cdk5 | G00000167 | SHANK1 | Q9WV48 | T635 |
| Cdk5 | G00000167 | Tau | P10637 | S493 |
| CK2 | G00000878 | Band 4.1like protein1 | Q9Z2H5 | S648 |
| CK2 | G00000878 | Band 4.1like protein1 | Q9Z2H5 | S650 |
| CK2 | G00000878 | cAMP-dependent protein kinase type II-alpha regulatory 1.E^E^E^Chain | P12367 | S75 |
| CK2 | G00000878 | cAMP-dependent protein kinase type II-alpha regulatory 1.E^E^E^Chain | P12367 | S77 |
| CK2 | G00000878 | cPLA2 | P47713 | S437 |
| CK2 | G00000878 | GSK3-beta | Q9WV60 | RGEPNVSYICSRYYR |
| CK2 | G00000878 | IRS1 | P35569 | S99 |
| CK2 | G00000878 | IRS1 | P35569 | SSEDLSNYASISFQK |
| CK2 | G00000878 | Neuromodulin | P06837 | AATKIQASFRGHITR |
| CK2 | G00000878 | NR2B | Q01097 | SSNGHVYEKLSSIES |
| CK2 | G00000878 | RasGap | P50904 | TVDGKEIYNTIRRKT |
| CKI | G00000169 | 14-3-3 zeta | P63101 | S184 |
| CKI | G00000169 | 14-3-3 zeta | P63101 | S57 |
| CKI | G00000169 | 14-3-3 zeta | P63101 | S58 |
| CKI | G00000169 | Band 4.1like protein1 | Q9Z2H5 | PELDRDKSDSETEGL |
| CKI | G00000169 | Bassoon | O88737 | S1417 |
| CKI | G00000169 | Bassoon | O88737 | S1509 |
| CKI | G00000169 | b-Catenin | Q02248 | QSYLDSGIHSGATTT |
| CKI | G00000169 | Calmodulin | P62204 | KDGDGTITTKELGTV |
| CKI | G00000169 | Calmodulin | P62204 | S81 |
| CKI | G00000169 | Calmodulin | P62204 | T79 |
| CKI | G00000169 | cAMP-dependent protein kinase type II-alpha regulatory 1.E^E^E^Chain | P12367 | S75 |
| CKI | G00000169 | cPLA2 | P47713 | LNTSYPLSPLRDFSS |
| CKI | G00000169 | cPLA2 | P47713 | RQNPSRCSVSLSNVE |
| CKI | G00000169 | d-Catenin-2 | O35927 | S531 |
| CKI | G00000169 | d-Catenin-2 | O35927 | S534 |
| CKI | G00000169 | DISKS large-associated protein 2 | Q8BJ42 | S450 |
| CKI | G00000169 | eNOS | P70313 | S1177 |
| CKI | G00000169 | GLUR2 | P23819 | NPSSSQNSQNFATYK |
| CKI | G00000169 | GLUR2 | P23819 | S863 |
| CKI | G00000169 | Guanine nucleotide-binding protein 3 | Q8BH66 | WGGFSEKSSDWSSEE |
| CKI | G00000169 | IRS1 | P35569 | PRSKSQSSSSCSNPI |
| CKI | G00000169 | IRS1 | P35569 | S105 |
| CKI | G00000169 | IRS1 | P35569 | S269 |
| CKI | G00000169 | IRS1 | P35569 | SSEDLSNYASISFQK |
| CKI | G00000169 | IRS1 | P35569 | VPNSRGDYMTMQIGC |
| CKI | G00000169 | MAP2 | P20357 | S1799 |
| CKI | G00000169 | MAP2 | P20357 | S1800 |
| CKI | G00000169 | MAP2 | P20357 | S1801 |
| CKI | G00000169 | mKIAA0942 protein | Q8CEA6 | TISRIGSTTNPFLDI |
| CKI | G00000169 | nArgbp2 | O35413 | KAEKRRGSLPDNSIL |
| CKI | G00000169 | Neuromodulin | P06837 | KATTDNSPSSKAED |
| CKI | G00000169 | nNOS | Q9Z0J4 | S1412 |
| CKI | G00000169 | nNOS | Q9Z0J4 | S847 |
| CKI | G00000169 | NR2A | P35436 | GRCPSDPYKHSLPSQ |
| CKI | G00000169 | NR2B | Q01097 | CKKAGNLYDISEDNS |
| CKI | G00000169 | NR2B | Q01097 | SSNGHVYEKLSSIES |
| CKI | G00000169 | Piccolo | Q9QYX7 | PTAVSLYSPTDEQSV |
| CKI | G00000169 | PKC-delta | P28867 | VQKKPTMYPEWKTTF |
| CKI | G00000169 | Plakophilin 4 | Q99569 | S512 |
| CKI | G00000169 | PLC-gamma1 | P19174 | EGSFESRYQQPFEDF |
| CKI | G00000169 | PLC-gamma1 | P19174 | KLAEGSAYEEVPTSM |
| CKI | G00000169 | Plexin 2 | P70207 | S1623 |
| CKI | G00000169 | PSD-95 | Q62108 | S418 |
| CKI | G00000169 | PTP-1B | P35821 | S70 |
| CKI | G00000169 | RAC1 | P63001 | S71 |
| CKI | G00000169 | RACK1 | P68040 | T228 |
| CKI | G00000169 | Rip11 | Q8R361 | THKRTYSDEASQLRA |
| CKI | G00000169 | S-adenosylhomocysteine hydrolase-like 1 | Q68FL4 | SSAASYTDSSDDETS |
| CKI | G00000169 | SAPAP3 | Q6PFD5 | S64 |
| CKI | G00000169 | SHANK1 | Q9WV48 | T635 |
| CKI | G00000169 | SHC | P29353 | TPPEELPSPSASSLGP |
| CKI | G00000169 | synGAP | Q9QUH6 | PSITKQHSQTPSTLN |
| CKI | G00000169 | synGAP | Q9QUH6 | S751 |
| CKI | G00000169 | synGAP | Q9QUH6 | S756 |
| CKI | G00000169 | Unc18-1 | O08599 | S146 |
| c-Raf | G00000185 | Calpain-1 | O35350 | S379 |
| c-Raf | G00000185 | Calpain-1 | O35350 | T380 |
| c-Raf | G00000185 | GLUR2 | P23819 | NPSSSQNSQNFATYK |
| c-Raf | G00000185 | IRS1 | P35569 | S105 |
| c-Raf | G00000185 | NR1 | P35438 | S889 |
| c-Raf | G00000185 | NR1 | P35438 | S890 |
| c-Raf | G00000185 | NR2B | Q01097 | S1303 |
| c-Raf | G00000185 | PDK-1 | Q9Z2A0 | S384 |
| c-Raf | G00000185 | PKC-delta | P28867 | EILQGLKYSFSVDWW |
| c-Raf | G00000185 | PKC-iota | Q62074 | S114 |
| c-Raf | G00000185 | PLC-gamma1 | P19174 | EGSFESRYQQPFEDF |
| c-Raf | G00000185 | SHANK1 | Q9WV48 | KRLFRHYTVGSYDSF |
| c-Raf | G00000185 | synGAP | Q9QUH6 | S750 |
| c-Raf | G00000185 | synGAP | Q9QUH6 | S751 |
| Erk1 | G00000179 | 14-3-3 zeta | P63101 | S57 |
| Erk1 | G00000179 | Band 4.1like protein1 | Q9Z2H5 | T475 |
| Erk1 | G00000179 | Bassoon | O88737 | KTLQRSLSDPKPLSP |
| Erk1 | G00000179 | Bassoon | O88737 | S3217 |
| Erk1 | G00000179 | Bassoon | O88737 | T2904 |
| Erk1 | G00000179 | Ca-Activated K Channel-alpha1 | Q08460 | S711 |
| Erk1 | G00000179 | CAMKII-beta | P28652 | T321 |
| Erk1 | G00000179 | Cortactin | Q60598 | RPPSSPIYEDAAPFK |
| Erk1 | G00000179 | cPLA2 | P47713 | S505 |
| Erk1 | G00000179 | DARPP32 | Q60829 | T34 |
| Erk1 | G00000179 | DISKS large-associated protein 2 | Q8BJ42 | S450 |
| Erk1 | G00000179 | Drebrin | Q9QXS6 | S140 |
| Erk1 | G00000179 | Drebrin | Q9QXS6 | S141 |
| Erk1 | G00000179 | GLUR2 | P23819 | S880 |
| Erk1 | G00000179 | IRS1 | P35569 | S612 |
| Erk1 | G00000179 | IRS1 | P35569 | S632 |
| Erk1 | G00000179 | MAP1B | P14873 | S1775 |
| Erk1 | G00000179 | MAP2 | P20357 | SMDEGDDYLPPTTPA |
| Erk1 | G00000179 | mGlur5 | P31424 | S1014 |
| Erk1 | G00000179 | mGlur5 | P31424 | S1016 |
| Erk1 | G00000179 | NR2A | P35436 | S1232 |
| Erk1 | G00000179 | Paralemmin | Q9Z0P4 | S157 |
| Erk1 | G00000179 | PKC-delta | P28867 | EILQGLKYSFSVDWW |
| Erk1 | G00000179 | Plexin 2 | P70207 | S1620 |
| Erk1 | G00000179 | PP1C-A | P62137 | T320 |
| Erk1 | G00000179 | PSD-95 | Q62108 | DYPTAMTPTSPRRYS |
| Erk1 | G00000179 | PSD-95 | Q62108 | S290 |
| Erk1 | G00000179 | PSD-95 | Q62108 | S295 |
| Erk1 | G00000179 | Rab3effector | Q9JIR4 | S895 |
| Erk1 | G00000179 | SAPAP3 | Q6PFD5 | S58 |
| Erk1 | G00000179 | SHC | P29353 | S36 |
| Erk1 | G00000179 | Src | P05480 | S73 |
| Erk1 | G00000179 | Synapsin1b | O88935 | S551 |
| Erk1 | G00000179 | Synapsin1b | O88935 | S62 |
| Erk1 | G00000179 | Synapsin1b | O88935 | S67 |
| Erk1 | G00000179 | Tau | P10637 | S493 |
| Erk1 | G00000179 | Tau | P10637 | T496 |
| Erk2 | G00000170 | 14-3-3 zeta | P63101 | S184 |
| Erk2 | G00000170 | 14-3-3 zeta | P63101 | S58 |
| Erk2 | G00000170 | Adenylate cyclase type IX | P51830 | S610 |
| Erk2 | G00000170 | Adenylate cyclase type IX | P51830 | S613 |
| Erk2 | G00000170 | Band 4.1like protein1 | Q9Z2H5 | T475 |
| Erk2 | G00000170 | Bassoon | O88737 | S1509 |
| Erk2 | G00000170 | Bassoon | O88737 | S3217 |
| Erk2 | G00000170 | Bassoon | O88737 | T2904 |
| Erk2 | G00000170 | b-Catenin | Q02248 | T653 |
| Erk2 | G00000170 | Ca-Activated K Channel-alpha1 | Q08460 | S711 |
| Erk2 | G00000170 | Calpain-1 | O35350 | S379 |
| Erk2 | G00000170 | Calpain-1 | O35350 | T380 |
| Erk2 | G00000170 | CAMKII-beta | P28652 | T321 |
| Erk2 | G00000170 | Cortactin | Q60598 | RPPSSPIYEDAAPFK |
| Erk2 | G00000170 | cPLA2 | P47713 | S505 |
| Erk2 | G00000170 | DARPP32 | Q60829 | T34 |
| Erk2 | G00000170 | DISKS large-associated protein 2 | Q8BJ42 | S450 |
| Erk2 | G00000170 | Drebrin | Q9QXS6 | S140 |
| Erk2 | G00000170 | Drebrin | Q9QXS6 | S141 |
| Erk2 | G00000170 | GLUR2 | P23819 | S880 |
| Erk2 | G00000170 | IRS1 | P35569 | S24 |
| Erk2 | G00000170 | IRS1 | P35569 | S612 |
| Erk2 | G00000170 | IRS1 | P35569 | S632 |
| Erk2 | G00000170 | IRS1 | P35569 | S887 |
| Erk2 | G00000170 | IRS1 | P35569 | TESITATSPASMVGG |
| Erk2 | G00000170 | MAP1B | P14873 | S1775 |
| Erk2 | G00000170 | MAP2 | P20357 | SMDEGDDYLPPTTPA |
| Erk2 | G00000170 | mGlur5 | P31424 | S1014 |
| Erk2 | G00000170 | mGlur5 | P31424 | S1016 |
| Erk2 | G00000170 | nNOS | Q9Z0J4 | S847 |
| Erk2 | G00000170 | NR2A | P35436 | S1232 |
| Erk2 | G00000170 | Paralemmin | Q9Z0P4 | S152 |
| Erk2 | G00000170 | Paralemmin | Q9Z0P4 | S157 |
| Erk2 | G00000170 | Paralemmin | Q9Z0P4 | T145 |
| Erk2 | G00000170 | PDK-1 | Q9Z2A0 | S384 |
| Erk2 | G00000170 | Piccolo | Q9QYX7 | S4260 |
| Erk2 | G00000170 | Piccolo | Q9QYX7 | S4263 |
| Erk2 | G00000170 | PKC-delta | P28867 | EILQGLKYSFSVDWW |
| Erk2 | G00000170 | PKC-iota | Q62074 | S114 |
| Erk2 | G00000170 | Plexin 2 | P70207 | S1620 |
| Erk2 | G00000170 | PP1C-A | P62137 | T320 |
| Erk2 | G00000170 | PSD-95 | Q62108 | DYPTAMTPTSPRRYS |
| Erk2 | G00000170 | PSD-95 | Q62108 | S290 |
| Erk2 | G00000170 | PSD-95 | Q62108 | S295 |
| Erk2 | G00000170 | PSD-95 | Q62108 | S35 |
| Erk2 | G00000170 | PSD-95 | Q62108 | T287 |
| Erk2 | G00000170 | PTP-1B | P35821 | ISEDVKSYYTVRQLE |
| Erk2 | G00000170 | PTP-1B | P35821 | S50 |
| Erk2 | G00000170 | Rab3effector | Q9JIR4 | S895 |
| Erk2 | G00000170 | SAPAP3 | Q6PFD5 | S58 |
| Erk2 | G00000170 | SHC | P29353 | S36 |
| Erk2 | G00000170 | Src | P05480 | S73 |
| Erk2 | G00000170 | Synapsin1b | O88935 | S551 |
| Erk2 | G00000170 | Tau | P10637 | S493 |
| Erk2 | G00000170 | Tau | P10637 | T496 |
| Erk2 | G00000170 | Voltage-gated potassium channel beta-2 subunit | P62482 | S20 |
| fes | G00000189 | 14-3-3 zeta | P63101 | FYYEILNSPEKACSL |
| fes | G00000189 | Bassoon | O88737 | Y1423 |
| fes | G00000189 | b-Catenin | Q02248 | Y654 |
| fes | G00000189 | b-Catenin | Q02248 | Y86 |
| fes | G00000189 | Calmodulin | P62204 | Y138 |
| fes | G00000189 | Calpain-1 | O35350 | Y387 |
| fes | G00000189 | Cortactin | Q60598 | Y421 |
| fes | G00000189 | Cortactin | Q60598 | Y466 |
| fes | G00000189 | GLUR1 | P23818 | GMIRVRKSKGKYAYL |
| fes | G00000189 | GLUR2 | P23819 | GVARVRKSKGKYAYL |
| fes | G00000189 | GLUR2 | P23819 | Y869 |
| fes | G00000189 | GLUR2 | P23819 | YKEGYNVYGIESVKI |
| fes | G00000189 | IRS1 | P35569 | Y1173 |
| fes | G00000189 | IRS1 | P35569 | Y1220 |
| fes | G00000189 | IRS1 | P35569 | Y460 |
| fes | G00000189 | IRS1 | P35569 | Y891 |
| fes | G00000189 | IRS1 | P35569 | Y983 |
| fes | G00000189 | NR2B | Q01097 | Y1304 |
| fes | G00000189 | NR2B | Q01097 | Y1336 |
| fes | G00000189 | NR2B | Q01097 | Y1472 |
| fes | G00000189 | PDK-1 | Q9Z2A0 | Y376 |
| fes | G00000189 | PDK-1 | Q9Z2A0 | Y379 |
| fes | G00000189 | PDK-1 | Q9Z2A0 | Y9 |
| fes | G00000189 | Pi3K-85 | P26450 | Y368 |
| fes | G00000189 | Pi3K-85 | P26450 | Y607 |
| fes | G00000189 | Pi3K-85 | P26450 | Y688 |
| fes | G00000189 | PKC-delta | P28867 | Y52 |
| fes | G00000189 | PKC-delta | P28867 | Y523 |
| fes | G00000189 | PKC-iota | Q62074 | Y116 |
| fes | G00000189 | PKC-iota | Q62074 | Y255 |
| fes | G00000189 | PLC-beta1 | Q9Z1B3 | Y334 |
| fes | G00000189 | PLC-gamma1 | P19174 | Y1253 |
| fes | G00000189 | PLC-gamma1 | P19174 | Y771 |
| fes | G00000189 | PLC-gamma1 | P19174 | Y775 |
| fes | G00000189 | PSD-95 | Q62108 | Y282 |
| fes | G00000189 | PTP-1B | P35821 | Y152 |
| fes | G00000189 | PTP-1B | P35821 | Y153 |
| fes | G00000189 | PTP-1B | P35821 | Y46 |
| fes | G00000189 | PTP-1B | P35821 | Y66 |
| fes | G00000189 | Pyk2 | Q9QVP9 | Y402 |
| fes | G00000189 | RAC1 | P63001 | YDRLRPLSYPQTDVF |
| fes | G00000189 | RACK1 | P68040 | Y227 |
| fes | G00000189 | RasGap | P50904 | Y451 |
| fes | G00000189 | SHANK1 | Q9WV48 | KRLFRHYTVGSYDSF |
| fes | G00000189 | SHC | P29353 | Y349 |
| fes | G00000189 | SHC | P29353 | Y350 |
| fes | G00000189 | Unc18-1 | O08599 | Y145 |
| fes | G00000189 | VAV | P27870 | Y174 |
| fes | G00000189 | Voltage-gated potassium channel beta-2 subunit | P62482 | Y25 |
| fyn | G00000186 | 14-3-3 zeta | P63101 | FYYEILNSPEKACSL |
| fyn | G00000186 | b-Catenin | Q02248 | Y654 |
| fyn | G00000186 | b-Catenin | Q02248 | Y86 |
| fyn | G00000186 | Calmodulin | P62204 | Y138 |
| fyn | G00000186 | Calpain-1 | O35350 | Y387 |
| fyn | G00000186 | Cortactin | Q60598 | Y421 |
| fyn | G00000186 | GLUR1 | P23818 | GMIRVRKSKGKYAYL |
| fyn | G00000186 | GLUR2 | P23819 | GVARVRKSKGKYAYL |
| fyn | G00000186 | GLUR2 | P23819 | Y869 |
| fyn | G00000186 | GLUR2 | P23819 | YKEGYNVYGIESVKI |
| fyn | G00000186 | IRS1 | P35569 | Y1220 |
| fyn | G00000186 | IRS1 | P35569 | Y983 |
| fyn | G00000186 | NR2B | Q01097 | Y1304 |
| fyn | G00000186 | NR2B | Q01097 | Y1336 |
| fyn | G00000186 | NR2B | Q01097 | Y1472 |
| fyn | G00000186 | PDK-1 | Q9Z2A0 | Y376 |
| fyn | G00000186 | PDK-1 | Q9Z2A0 | Y379 |
| fyn | G00000186 | Pi3K-85 | P26450 | Y607 |
| fyn | G00000186 | Pi3K-85 | P26450 | Y688 |
| fyn | G00000186 | PKC-iota | Q62074 | Y116 |
| fyn | G00000186 | PKC-iota | Q62074 | Y270 |
| fyn | G00000186 | PLC-gamma1 | P19174 | Y1253 |
| fyn | G00000186 | PLC-gamma1 | P19174 | Y771 |
| fyn | G00000186 | PLC-gamma1 | P19174 | Y775 |
| fyn | G00000186 | PSD-95 | Q62108 | Y439 |
| fyn | G00000186 | PTP-1B | P35821 | Y66 |
| fyn | G00000186 | Pyk2 | Q9QVP9 | Y402 |
| fyn | G00000186 | RACK1 | P68040 | Y227 |
| fyn | G00000186 | RasGap | P50904 | Y451 |
| fyn | G00000186 | SHANK1 | Q9WV48 | KRLFRHYTVGSYDSF |
| fyn | G00000186 | SHC | P29353 | Y349 |
| fyn | G00000186 | SHC | P29353 | Y350 |
| fyn | G00000186 | VAV | P27870 | Y174 |
| fyn | G00000186 | Voltage-gated potassium channel beta-2 subunit | P62482 | Y25 |
| GSK3-beta | G00000171 | Adenylate cyclase type IX | P51830 | S610 |
| GSK3-beta | G00000171 | Adenylate cyclase type IX | P51830 | S613 |
| GSK3-beta | G00000171 | Bassoon | O88737 | PRATAEFSTQTPSLT |
| GSK3-beta | G00000171 | Bassoon | O88737 | S3217 |
| GSK3-beta | G00000171 | b-Catenin | Q02248 | QSYLDSGIHSGATTT |
| GSK3-beta | G00000171 | b-Catenin | Q02248 | S33 |
| GSK3-beta | G00000171 | Ca-Activated K Channel-alpha1 | Q08460 | S923 |
| GSK3-beta | G00000171 | Ca-Activated K Channel-alpha1 | Q08460 | S924 |
| GSK3-beta | G00000171 | Calmodulin | P62204 | T44 |
| GSK3-beta | G00000171 | cPLA2 | P47713 | S505 |
| GSK3-beta | G00000171 | GLUR2 | P23819 | S880 |
| GSK3-beta | G00000171 | GLUR4 | Q9Z2W8 | S862 |
| GSK3-beta | G00000171 | IRS1 | P35569 | NLHTDDGYMPMSPGV |
| GSK3-beta | G00000171 | IRS1 | P35569 | S632 |
| GSK3-beta | G00000171 | IRS1 | P35569 | S887 |
| GSK3-beta | G00000171 | MAP2 | P20357 | SMDEGDDYLPPTTPA |
| GSK3-beta | G00000171 | mGlur3 | Q9QYS2 | S845 |
| GSK3-beta | G00000171 | Paralemmin | Q9Z0P4 | S157 |
| GSK3-beta | G00000171 | Plexin 2 | P70207 | S1620 |
| GSK3-beta | G00000171 | PSD-95 | Q62108 | S290 |
| GSK3-beta | G00000171 | PSD-95 | Q62108 | S418 |
| GSK3-beta | G00000171 | PSD-95 | Q62108 | T19 |
| GSK3-beta | G00000171 | PTP-1B | P35821 | S55 |
| GSK3-beta | G00000171 | SAPAP3 | Q6PFD5 | S58 |
| GSK3-beta | G00000171 | SHANK1 | Q9WV48 | T635 |
| GSK3-beta | G00000171 | Src | P05480 | S73 |
| GSK3-beta | G00000171 | Stargazin | O88602 | T320 |
| GSK3-beta | G00000171 | Stargazin | O88602 | T321 |
| GSK3-beta | G00000171 | Tau | P10637 | S486 |
| GSK3-beta | G00000171 | Tau | P10637 | S490 |
| GSK3-beta | G00000171 | Unc18-1 | O08599 | S142 |
| GSK3-beta | G00000171 | Voltage-gated potassium channel beta-2 subunit | P62482 | S20 |
| JNK3 | G00000887 | cPLA2 | P47713 | S505 |
| JNK3 | G00000887 | DARPP32 | Q60829 | T34 |
| JNK3 | G00000887 | Drebrin | Q9QXS6 | S140 |
| JNK3 | G00000887 | GSK3-beta | Q9WV60 | RGEPNVSYICSRYYR |
| JNK3 | G00000887 | PKC-delta | P28867 | EILQGLKYSFSVDWW |
| JNK3 | G00000887 | PSD-95 | Q62108 | T287 |
| JNK3 | G00000887 | Rab3effector | Q9JIR4 | S895 |
| JNK3 | G00000887 | SHANK1 | Q9WV48 | KRLFRHYTVGSYDSF |
| MEK1 | G00000173 | Calpain-1 | O35350 | S379 |
| MEK1 | G00000173 | Calpain-1 | O35350 | T380 |
| MEK1 | G00000173 | Drebrin | Q9QXS6 | S140 |
| MEK1 | G00000173 | Drebrin | Q9QXS6 | S141 |
| MEK1 | G00000173 | GLUR1 | P23818 | GMIRVRKSKGKYAYL |
| MEK1 | G00000173 | GLUR2 | P23819 | GVARVRKSKGKYAYL |
| MEK1 | G00000173 | GLUR2 | P23819 | Y869 |
| MEK1 | G00000173 | IRS1 | P35569 | S24 |
| MEK1 | G00000173 | Neurofascin precursor | Q810U3 | ETEGNESSEATSPVN |
| MEK1 | G00000173 | NR1 | P35438 | S889 |
| MEK1 | G00000173 | PKC-delta | P28867 | VQKKPTMYPEWKTTF |
| MEK1 | G00000173 | PKC-iota | Q62074 | S114 |
| MEK1 | G00000173 | PKC-iota | Q62074 | S254 |
| MEK1 | G00000173 | PKC-iota | Q62074 | T266 |
| MEK1 | G00000173 | PKC-iota | Q62074 | Y255 |
| MEK1 | G00000173 | SHANK1 | Q9WV48 | KRLFRHYTVGSYDSF |
| p38-alpha | Band 4.1like protein1 | Q9Z2H5 | T475 | |
| p38-alpha | Bassoon | O88737 | S2865 | |
| p38-alpha | Bassoon | O88737 | S3217 | |
| p38-alpha | Bassoon | O88737 | T2904 | |
| p38-alpha | Calmodulin | P62204 | KDGDGTITTKELGTV | |
| p38-alpha | Calpain-1 | O35350 | S379 | |
| p38-alpha | Calpain-1 | O35350 | T380 | |
| p38-alpha | Cortactin | Q60598 | RPPSSPIYEDAAPFK | |
| p38-alpha | cPLA2 | P47713 | S505 | |
| p38-alpha | DARPP32 | Q60829 | T34 | |
| p38-alpha | DISKS large-associated protein 2 | Q8BJ42 | S450 | |
| p38-alpha | Drebrin | Q9QXS6 | S140 | |
| p38-alpha | Drebrin | Q9QXS6 | S141 | |
| p38-alpha | GLUR2 | P23819 | S880 | |
| p38-alpha | IRS1 | P35569 | S612 | |
| p38-alpha | IRS1 | P35569 | S632 | |
| p38-alpha | IRS1 | P35569 | TESITATSPASMVGG | |
| p38-alpha | MAP1B | P14873 | S1775 | |
| p38-alpha | MAP2 | P20357 | SMDEGDDYLPPTTPA | |
| p38-alpha | mGlur4a | P31423 | S859 | |
| p38-alpha | mGlur4a | P31423 | T865 | |
| p38-alpha | mGlur5 | P31424 | S1014 | |
| p38-alpha | mGlur5 | P31424 | S1016 | |
| p38-alpha | mGlur7a | Q68ED2 | T868 | |
| p38-alpha | Paralemmin | Q9Z0P4 | T141 | |
| p38-alpha | PKC-delta | P28867 | EILQGLKYSFSVDWW | |
| p38-alpha | Plexin 2 | P70207 | S1620 | |
| p38-alpha | PSD-95 | Q62108 | LMNSSLGSGTASLRS | |
| p38-alpha | PSD-95 | Q62108 | S290 | |
| p38-alpha | PSD-95 | Q62108 | T287 | |
| p38-alpha | Rab3effector | Q9JIR4 | S895 | |
| p38-alpha | SAPAP3 | Q6PFD5 | S58 | |
| p38-alpha | Src | P05480 | S73 | |
| p38-alpha | Synapsin1b | O88935 | S551 | |
| p38-alpha | Tau | P10637 | S490 | |
| p38-alpha | Tau | P10637 | SGYSSPGSPGTPGSR | |
| p70S6K | G00000177 | 14-3-3 zeta | P63101 | S58 |
| p70S6K | G00000177 | Band 4.1like protein1 | Q9Z2H5 | S466 |
| p70S6K | G00000177 | Bassoon | O88737 | S1418 |
| p70S6K | G00000177 | Bassoon | O88737 | T1510 |
| p70S6K | G00000177 | Calpain-1 | O35350 | S379 |
| p70S6K | G00000177 | Calpain-1 | O35350 | T380 |
| p70S6K | G00000177 | cPLA2 | P47713 | S726 |
| p70S6K | G00000177 | Drebrin | Q9QXS6 | S141 |
| p70S6K | G00000177 | eNOS | P70313 | S1177 |
| p70S6K | G00000177 | eNOS | P70313 | S613 |
| p70S6K | G00000177 | eNOS | P70313 | S631 |
| p70S6K | G00000177 | GLUR1 | P23818 | T858 |
| p70S6K | G00000177 | IRS1 | P35569 | PRSKSQSSSSCSNPI |
| p70S6K | G00000177 | IRS1 | P35569 | S267 |
| p70S6K | G00000177 | IRS1 | P35569 | S302 |
| p70S6K | G00000177 | IRS1 | P35569 | SSEDLSNYASISFQK |
| p70S6K | G00000177 | MAP2 | P20357 | S1799 |
| p70S6K | G00000177 | MAP2 | P20357 | S1800 |
| p70S6K | G00000177 | mGlur4a | P31423 | S859 |
| p70S6K | G00000177 | mGlur7a | Q68ED2 | S862 |
| p70S6K | G00000177 | nNOS | Q9Z0J4 | SYKVRFNSVSSYSDS |
| p70S6K | G00000177 | NR1 | P35438 | S896 |
| p70S6K | G00000177 | NR1 | P35438 | S897 |
| p70S6K | G00000177 | NR2A | P35436 | GRCPSDPYKHSLPSQ |
| p70S6K | G00000177 | PSD-95 | Q62108 | DYPTAMTPTSPRRYS |
| p70S6K | G00000177 | RasGap | P50904 | TVDGKEIYNTIRRKT |
| p70S6K | G00000177 | Rip11 | Q8R361 | S307 |
| p70S6K | G00000177 | SHANK1 | Q9WV48 | KRLFRHYTVGSYDSF |
| p70S6K | G00000177 | Synapsin1b | O88935 | S9 |
| p70S6K | G00000177 | Voltage-gated potassium channel beta-2 subunit | P62482 | LSLRQTGSPGMIYST |
| PDK-1 | G00000184 | GLUR1 | P23818 | S863 |
| PDK-1 | G00000184 | PKC-delta | P28867 | T50 |
| PDK-1 | G00000184 | PKC-delta | P28867 | VQKKPTMYPEWKTTF |
| PDK-1 | G00000184 | SHANK1 | Q9WV48 | KRLFRHYTVGSYDSF |
| PDK-1 | G00000184 | synGAP | Q9QUH6 | S750 |
| PKA | G00001000 | 14-3-3 zeta | P63101 | S58 |
| PKA | G00001000 | Bassoon | O88737 | S2121 |
| PKA | G00001000 | Bassoon | O88737 | S3213 |
| PKA | G00001000 | b-Catenin | Q02248 | T653 |
| PKA | G00001000 | Calpain-1 | O35350 | GTWRRGSTAGGCRNY |
| PKA | G00001000 | Calpain-1 | O35350 | S379 |
| PKA | G00001000 | CAMKII-beta | P28652 | S315 |
| PKA | G00001000 | cPLA2 | P47713 | LNTSYPLSPLRDFSS |
| PKA | G00001000 | DARPP32 | Q60829 | T34 |
| PKA | G00001000 | d-Catenin-2 | O35927 | S531 |
| PKA | G00001000 | DISKS large-associated protein 2 | Q8BJ42 | S440 |
| PKA | G00001000 | Drebrin | Q9QXS6 | S140 |
| PKA | G00001000 | GABA-B R2 | O88871 | S892 |
| PKA | G00001000 | IRS1 | P35569 | S265 |
| PKA | G00001000 | IRS1 | P35569 | SSEDLSNYASISFQK |
| PKA | G00001000 | MBP | P04370 | S144 |
| PKA | G00001000 | mGlur2 | P31421 | S843 |
| PKA | G00001000 | mGlur4a | P31423 | S859 |
| PKA | G00001000 | mGlur7a | Q68ED2 | S862 |
| PKA | G00001000 | mKIAA0942 protein | Q8CEA6 | S46 |
| PKA | G00001000 | nArgbp2 | O35413 | S844 |
| PKA | G00001000 | NR1 | P35438 | S897 |
| PKA | G00001000 | NR2A | P35436 | GRCPSDPYKHSLPSQ |
| PKA | G00001000 | NR2B | Q01097 | S1303 |
| PKA | G00001000 | PKC-delta | P28867 | EILQGLKYSFSVDWW |
| PKA | G00001000 | PKC-iota | Q62074 | S254 |
| PKA | G00001000 | PSD-95 | Q62108 | DYPTAMTPTSPRRYS |
| PKA | G00001000 | RAP1B | Q99JI6 | S179 |
| PKA | G00001000 | RasGap | P50904 | TVDGKEIYNTIRRKT |
| PKA | G00001000 | SHANK1 | Q9WV48 | T635 |
| PKA | G00001000 | Synapsin1b | O88935 | S9 |
| PKA | G00001000 | synGAP | Q9QUH6 | S750 |
| PKA | G00001000 | synGAP | Q9QUH6 | S751 |
| PKC-alpha | G00000159 | 14-3-3 zeta | P63101 | S58 |
| PKC-alpha | G00000159 | Ankyrin2 | Q01484 | S31 |
| PKC-alpha | G00000159 | Ankyrin2 | Q01484 | S34 |
| PKC-alpha | G00000159 | Calpain-1 | O35350 | S379 |
| PKC-alpha | G00000159 | Calpain-1 | O35350 | T380 |
| PKC-alpha | G00000159 | DARPP32 | Q60829 | T34 |
| PKC-alpha | G00000159 | d-Catenin-2 | O35927 | S531 |
| PKC-alpha | G00000159 | DISKS large-associated protein 2 | Q8BJ42 | S440 |
| PKC-alpha | G00000159 | Drebrin | Q9QXS6 | S140 |
| PKC-alpha | G00000159 | GLUR1 | P23818 | S710 |
| PKC-alpha | G00000159 | GLUR2 | P23819 | S717 |
| PKC-alpha | G00000159 | Guanine nucleotide-binding protein 3 | Q8BH66 | WGGFSEKSSDWSSEE |
| PKC-alpha | G00000159 | IRS1 | P35569 | S24 |
| PKC-alpha | G00000159 | IRS1 | P35569 | SSEDLSNYASISFQK |
| PKC-alpha | G00000159 | MAP2 | P20357 | S1800 |
| PKC-alpha | G00000159 | MBP | P04370 | S144 |
| PKC-alpha | G00000159 | mGlur4a | P31423 | S859 |
| PKC-alpha | G00000159 | mGlur7a | Q68ED2 | S862 |
| PKC-alpha | G00000159 | mKIAA0942 protein | Q8CEA6 | S46 |
| PKC-alpha | G00000159 | NR1 | P35438 | S889 |
| PKC-alpha | G00000159 | NR1 | P35438 | S890 |
| PKC-alpha | G00000159 | PKC-delta | P28867 | EILQGLKYSFSVDWW |
| PKC-alpha | G00000159 | PKC-iota | Q62074 | S254 |
| PKC-alpha | G00000159 | PP1C-A | P62137 | T320 |
| PKC-alpha | G00000159 | PSD-95 | Q62108 | DYPTAMTPTSPRRYS |
| PKC-alpha | G00000159 | RAP1B | Q99JI6 | S179 |
| PKC-alpha | G00000159 | RasGap | P50904 | TVDGKEIYNTIRRKT |
| PKC-alpha | G00000159 | SHANK1 | Q9WV48 | T635 |
| PKC-alpha | G00000159 | synGAP | Q9QUH6 | S750 |
| PKC-alpha | G00000159 | synGAP | Q9QUH6 | S751 |
| PKC-delta | 14-3-3 zeta | P63101 | S57 | |
| PKC-delta | 14-3-3 zeta | P63101 | S58 | |
| PKC-delta | Ankyrin2 | Q01484 | S31 | |
| PKC-delta | Ankyrin2 | Q01484 | S34 | |
| PKC-delta | Ca-Activated K Channel-alpha1 | Q08460 | T709 | |
| PKC-delta | Calpain-1 | O35350 | S379 | |
| PKC-delta | Calpain-1 | O35350 | T380 | |
| PKC-delta | DARPP32 | Q60829 | T34 | |
| PKC-delta | Drebrin | Q9QXS6 | S140 | |
| PKC-delta | GLUR1 | P23818 | S710 | |
| PKC-delta | GLUR2 | P23819 | S717 | |
| PKC-delta | IRS1 | P35569 | S24 | |
| PKC-delta | IRS1 | P35569 | SSEDLSNYASISFQK | |
| PKC-delta | mGlur4a | P31423 | S859 | |
| PKC-delta | mGlur7a | Q68ED2 | S862 | |
| PKC-delta | NR1 | P35438 | S889 | |
| PKC-delta | NR1 | P35438 | S890 | |
| PKC-delta | PKC-delta | P28867 | EILQGLKYSFSVDWW | |
| PKC-delta | PKC-iota | Q62074 | S254 | |
| PKC-delta | PP1C-A | P62137 | T320 | |
| PKC-delta | PSD-95 | Q62108 | DYPTAMTPTSPRRYS | |
| PKC-delta | RAP1A | P62835 | S180 | |
| PKC-delta | RAP1B | Q99JI6 | S179 | |
| PKC-delta | RasGap | P50904 | TVDGKEIYNTIRRKT | |
| PKC-delta | SHANK1 | Q9WV48 | KRLFRHYTVGSYDSF | |
| PKC-zeta | 14-3-3 zeta | P63101 | S57 | |
| PKC-zeta | 14-3-3 zeta | P63101 | S58 | |
| PKC-zeta | Ankyrin2 | Q01484 | S34 | |
| PKC-zeta | Calpain-1 | O35350 | S379 | |
| PKC-zeta | Calpain-1 | O35350 | T380 | |
| PKC-zeta | Drebrin | Q9QXS6 | S140 | |
| PKC-zeta | GABA-B R2 | O88871 | S892 | |
| PKC-zeta | GLUR1 | P23818 | S710 | |
| PKC-zeta | GLUR2 | P23819 | S717 | |
| PKC-zeta | IRS1 | P35569 | S24 | |
| PKC-zeta | IRS1 | P35569 | SSEDLSNYASISFQK | |
| PKC-zeta | mGlur4a | P31423 | S859 | |
| PKC-zeta | mGlur7a | Q68ED2 | S862 | |
| PKC-zeta | NR1 | P35438 | S889 | |
| PKC-zeta | NR1 | P35438 | S890 | |
| PKC-zeta | PKC-iota | Q62074 | S254 | |
| PKC-zeta | PKC-iota | Q62074 | T266 | |
| PKC-zeta | Plexin 2 | P70207 | SFRYTGSPDSLRSRV | |
| PKC-zeta | PP1C-A | P62137 | T320 | |
| PKC-zeta | PSD-95 | Q62108 | DYPTAMTPTSPRRYS | |
| PKC-zeta | PSD-95 | Q62108 | LMNSSLGSGTASLRS | |
| PKC-zeta | PTP-1B | P35821 | S55 | |
| PKC-zeta | RAP1B | Q99JI6 | S179 | |
| PKC-zeta | RasGap | P50904 | TVDGKEIYNTIRRKT | |
| PKC-zeta | Synapsin1b | O88935 | S9 | |
| Pyk2 | G00000051 | 14-3-3 zeta | P63101 | FYYEILNSPEKACSL |
| Pyk2 | G00000051 | Bassoon | O88737 | Y1423 |
| Pyk2 | G00000051 | b-Catenin | Q02248 | Y654 |
| Pyk2 | G00000051 | b-Catenin | Q02248 | Y86 |
| Pyk2 | G00000051 | Calmodulin | P62204 | Y138 |
| Pyk2 | G00000051 | Calpain-1 | O35350 | Y387 |
| Pyk2 | G00000051 | Cortactin | Q60598 | Y421 |
| Pyk2 | G00000051 | GLUR1 | P23818 | GMIRVRKSKGKYAYL |
| Pyk2 | G00000051 | GLUR2 | P23819 | GVARVRKSKGKYAYL |
| Pyk2 | G00000051 | GLUR2 | P23819 | Y869 |
| Pyk2 | G00000051 | GLUR2 | P23819 | YKEGYNVYGIESVKI |
| Pyk2 | G00000051 | IRS1 | P35569 | Y1173 |
| Pyk2 | G00000051 | IRS1 | P35569 | Y1220 |
| Pyk2 | G00000051 | IRS1 | P35569 | Y983 |
| Pyk2 | G00000051 | NR2B | Q01097 | Y1304 |
| Pyk2 | G00000051 | NR2B | Q01097 | Y1336 |
| Pyk2 | G00000051 | NR2B | Q01097 | Y1472 |
| Pyk2 | G00000051 | PDK-1 | Q9Z2A0 | Y376 |
| Pyk2 | G00000051 | PDK-1 | Q9Z2A0 | Y379 |
| Pyk2 | G00000051 | PDK-1 | Q9Z2A0 | Y9 |
| Pyk2 | G00000051 | Pi3K-85 | P26450 | Y607 |
| Pyk2 | G00000051 | Pi3K-85 | P26450 | Y688 |
| Pyk2 | G00000051 | Piccolo | Q9QYX7 | Y3867 |
| Pyk2 | G00000051 | PLC-gamma1 | P19174 | Y1253 |
| Pyk2 | G00000051 | PLC-gamma1 | P19174 | Y771 |
| Pyk2 | G00000051 | PLC-gamma1 | P19174 | Y775 |
| Pyk2 | G00000051 | PSD-95 | Q62108 | Y282 |
| Pyk2 | G00000051 | PTP-1B | P35821 | Y66 |
| Pyk2 | G00000051 | RACK1 | P68040 | Y227 |
| Pyk2 | G00000051 | RasGap | P50904 | Y451 |
| Pyk2 | G00000051 | SHC | P29353 | Y349 |
| Pyk2 | G00000051 | SHC | P29353 | Y350 |
| Pyk2 | G00000051 | VAV | P27870 | Y174 |
| Pyk2 | G00000051 | Voltage-gated potassium channel beta-2 subunit | P62482 | Y25 |
| Rock2 | G00000163 | Calpain-1 | O35350 | S379 |
| Rock2 | G00000163 | Calpain-1 | O35350 | T380 |
| Rock2 | G00000163 | DARPP32 | Q60829 | T34 |
| Rock2 | G00000163 | GLUR2 | P23819 | S717 |
| Rock2 | G00000163 | IRS1 | P35569 | NLHTDDGYMPMSPGV |
| Rock2 | G00000163 | IRS1 | P35569 | S24 |
| Rock2 | G00000163 | mGlur4a | P31423 | S859 |
| Rock2 | G00000163 | mGlur7a | Q68ED2 | S862 |
| Rock2 | G00000163 | nNOS | Q9Z0J4 | S847 |
| Rock2 | G00000163 | NR1 | P35438 | S889 |
| Rock2 | G00000163 | NR1 | P35438 | S890 |
| Rock2 | G00000163 | PKC-iota | Q62074 | T266 |
| Rock2 | G00000163 | PSD-95 | Q62108 | DYPTAMTPTSPRRYS |
| Rock2 | G00000163 | RAP1A | P62835 | S180 |
| Rock2 | G00000163 | RAP1B | Q99JI6 | S179 |
| Rock2 | G00000163 | RasGap | P50904 | TVDGKEIYNTIRRKT |
| Rock2 | G00000163 | Tau | P10637 | SGYSSPGSPGTPGSR |
| RSK-2 | G00000178 | 14-3-3 zeta | P63101 | VVGARRSSWRVVSSI |
| RSK-2 | G00000178 | Ankyrin2 | Q01484 | S31 |
| RSK-2 | G00000178 | Bassoon | O88737 | S1415 |
| RSK-2 | G00000178 | Bassoon | O88737 | S1417 |
| RSK-2 | G00000178 | Calpain-1 | O35350 | S379 |
| RSK-2 | G00000178 | Calpain-1 | O35350 | T380 |
| RSK-2 | G00000178 | eNOS | P70313 | S1177 |
| RSK-2 | G00000178 | eNOS | P70313 | S613 |
| RSK-2 | G00000178 | GLUR2 | P23819 | S717 |
| RSK-2 | G00000178 | IRS1 | P35569 | S265 |
| RSK-2 | G00000178 | IRS1 | P35569 | SSEDLSNYASISFQK |
| RSK-2 | G00000178 | MAP2 | P20357 | S1799 |
| RSK-2 | G00000178 | MAP2 | P20357 | S1800 |
| RSK-2 | G00000178 | mKIAA0942 protein | Q8CEA6 | T48 |
| RSK-2 | G00000178 | nArgbp2 | O35413 | S844 |
| RSK-2 | G00000178 | nNOS | Q9Z0J4 | SYKVRFNSVSSYSDS |
| RSK-2 | G00000178 | NR2A | P35436 | GRCPSDPYKHSLPSQ |
| RSK-2 | G00000178 | NR2B | Q01097 | S1303 |
| RSK-2 | G00000178 | PKC-delta | P28867 | EILQGLKYSFSVDWW |
| RSK-2 | G00000178 | PLC-gamma1 | P19174 | EGSFESRYQQPFEDF |
| RSK-2 | G00000178 | PSD-95 | Q62108 | DYPTAMTPTSPRRYS |
| RSK-2 | G00000178 | PTP-1B | P35821 | S50 |
| RSK-2 | G00000178 | PTP-1B | P35821 | S70 |
| RSK-2 | G00000178 | RAP1B | Q99JI6 | S179 |
| RSK-2 | G00000178 | Rip11 | Q8R361 | S307 |
| RSK-2 | G00000178 | SHANK1 | Q9WV48 | KRLFRHYTVGSYDSF |
| RSK-2 | G00000178 | Synapsin1b | O88935 | S9 |
| RSK-2 | G00000178 | synGAP | Q9QUH6 | S751 |
| RSK-2 | G00000178 | Voltage-gated potassium channel beta-2 subunit | P62482 | LSLRQTGSPGMIYST |
| Src | G00000888 | 14-3-3 zeta | P63101 | FYYEILNSPEKACSL |
| Src | G00000888 | Bassoon | O88737 | Y1423 |
| Src | G00000888 | b-Catenin | Q02248 | Y654 |
| Src | G00000888 | b-Catenin | Q02248 | Y86 |
| Src | G00000888 | Calmodulin | P62204 | Y138 |
| Src | G00000888 | Calpain-1 | O35350 | Y387 |
| Src | G00000888 | Cortactin | Q60598 | Y421 |
| Src | G00000888 | GLUR1 | P23818 | GMIRVRKSKGKYAYL |
| Src | G00000888 | GLUR2 | P23819 | GVARVRKSKGKYAYL |
| Src | G00000888 | GLUR2 | P23819 | Y869 |
| Src | G00000888 | GLUR2 | P23819 | YKEGYNVYGIESVKI |
| Src | G00000888 | IRS1 | P35569 | Y1220 |
| Src | G00000888 | IRS1 | P35569 | Y891 |
| Src | G00000888 | IRS1 | P35569 | Y983 |
| Src | G00000888 | NR2B | Q01097 | Y1304 |
| Src | G00000888 | NR2B | Q01097 | Y1336 |
| Src | G00000888 | NR2B | Q01097 | Y1472 |
| Src | G00000888 | PDK-1 | Q9Z2A0 | Y376 |
| Src | G00000888 | PDK-1 | Q9Z2A0 | Y379 |
| Src | G00000888 | Pi3K-85 | P26450 | Y688 |
| Src | G00000888 | PKC-iota | Q62074 | Y116 |
| Src | G00000888 | PKC-iota | Q62074 | Y270 |
| Src | G00000888 | PLC-gamma1 | P19174 | Y1253 |
| Src | G00000888 | PLC-gamma1 | P19174 | Y771 |
| Src | G00000888 | PLC-gamma1 | P19174 | Y775 |
| Src | G00000888 | PTP-1B | P35821 | Y66 |
| Src | G00000888 | Pyk2 | Q9QVP9 | Y402 |
| Src | G00000888 | RACK1 | P68040 | Y227 |
| Src | G00000888 | RasGap | P50904 | Y451 |
| Src | G00000888 | SHANK1 | Q9WV48 | KRLFRHYTVGSYDSF |
| Src | G00000888 | SHC | P29353 | Y349 |
| Src | G00000888 | SHC | P29353 | Y350 |
| Src | G00000888 | VAV | P27870 | Y174 |
| Src | G00000888 | Voltage-gated potassium channel beta-2 subunit | P62482 | Y25 |
| TBKI | G00000164 | 14-3-3 zeta | P63101 | S184 |
| TBKI | G00000164 | 14-3-3 zeta | P63101 | S58 |
| TBKI | G00000164 | Adenylate cyclase type IX | P51830 | RGQGTASPGSVSDLA |
| TBKI | G00000164 | Ankyrin2 | Q01484 | S34 |
| TBKI | G00000164 | Bassoon | O88737 | ERSPSTSSTIHSYGQ |
| TBKI | G00000164 | Bassoon | O88737 | PRATAEFSTQTPSLT |
| TBKI | G00000164 | Bassoon | O88737 | S3217 |
| TBKI | G00000164 | Calpain-m (2) | O08529 | GALFQDPSFPALPSS |
| TBKI | G00000164 | cPLA2 | P47713 | LNTSYPLSPLRDFSS |
| TBKI | G00000164 | d-Catenin-2 | O35927 | S534 |
| TBKI | G00000164 | IRS1 | P35569 | S887 |
| TBKI | G00000164 | IRS1 | P35569 | TESITATSPASMVGG |
| TBKI | G00000164 | MAP2 | P20357 | S1161 |
| TBKI | G00000164 | MAP2 | P20357 | S1801 |
| TBKI | G00000164 | mGlur5 | P31424 | AAARPRSPSPISTLS |
| TBKI | G00000164 | mKIAA0942 protein | Q8CEA6 | TISRIGSTTNPFLDI |
| TBKI | G00000164 | nArgbp2 | O35413 | S844 |
| TBKI | G00000164 | nArgbp2 | O35413 | S849 |
| TBKI | G00000164 | nNOS | Q9Z0J4 | SYKVRFNSVSSYSDS |
| TBKI | G00000164 | NR2B | Q01097 | SSNGHVYEKLSSIES |
| TBKI | G00000164 | NR2B | Q01097 | T1306 |
| TBKI | G00000164 | PDK-1 | Q9Z2A0 | S384 |
| TBKI | G00000164 | Pi3K-85 | P26450 | FAEPYNLYSSLKELV |
| TBKI | G00000164 | Piccolo | Q9QYX7 | AQNSEEESPLSPVGQ |
| TBKI | G00000164 | PLC-gamma1 | P19174 | KLAEGSAYEEVPTSM |
| TBKI | G00000164 | Plexin 2 | P70207 | S1623 |
| TBKI | G00000164 | PSD-95 | Q62108 | DYPTAMTPTSPRRYS |
| TBKI | G00000164 | PSD-95 | Q62108 | S415 |
| TBKI | G00000164 | PSD-95 | Q62108 | T287 |
| TBKI | G00000164 | PTP-1B | P35821 | S50 |
| TBKI | G00000164 | PTP-1B | P35821 | S55 |
| TBKI | G00000164 | SAPAP3 | Q6PFD5 | S58 |
| TBKI | G00000164 | SAPAP3 | Q6PFD5 | S64 |
| TBKI | G00000164 | SHANK1 | Q9WV48 | KRLFRHYTVGSYDSF |
| TBKI | G00000164 | SHC | P29353 | TPPEELPSPSASSLGP |
| TBKI | G00000164 | synGAP | Q9QUH6 | PSITKQHSQTPSTLN |
| TBKI | G00000164 | synGAP | Q9QUH6 | S750 |
| TBKI | G00000164 | synGAP | Q9QUH6 | S756 |
| TBKI | G00000164 | synGAP | Q9QUH6 | S765 |
| TBKI | G00000164 | Tau | P10637 | SGYSSPGSPGTPGSR |
| TBKI | G00000164 | Unc18-1 | O08599 | S146 |
| TBKI | G00000164 | Unc18-1 | O08599 | S149 |
| TBKI | G00000164 | Voltage-gated potassium channel beta-2 subunit | P62482 | LSLRQTGSPGMIYST |
Table 3. Neurotransmitter receptor modulation of synaptic protein phosphorylation
The table reports the phosphorylation and dephosphorylation that occurs in protein substrates isolated from the postsynaptic density of hippocampus slices stimulated with NMDA (at 3 minutes).
| Gene | G2Cb gene | UniProt | Family | Sub-family | NMDA effect |
|---|---|---|---|---|---|
| Ctnnd1 | G00000555 | P30999 | Cell Adhesion | Catenins | dephosphorylated |
| Cadm4 | Q8R464 | Cell Adhesion | Other Cell Adhesion Molecules | dephosphorylated | |
| Col27a1 | Q5QNQ9 | Cell Adhesion | Other Cell Adhesion Molecules | dephosphorylated | |
| Jph1 | Q9ET80 | Cell Adhesion | Other Cell Adhesion Molecules | phosphorylated | |
| Pkp2 | Q3TIY5 | Cell Adhesion | Other Cell Adhesion Molecules | dephosphorylated | |
| Pkp4 | G00000595 | Q68FH0 | Cell Adhesion | Other Cell Adhesion Molecules | phosphorylated + dephosphorylated |
| Aprin | Q4VA53 | Cell cycle | Cell cycle | dephosphorylated | |
| Tbrg1 | Q3UB74 | Cell cycle | Cell cycle | dephosphorylated | |
| Gria2 | G00000058 | P23819 | Channels and Receptors | Glutamate Receptors | dephosphorylated |
| Grm5 | G00000064 | P31424 | Channels and Receptors | Glutamate Receptors | dephosphorylated |
| Atp8a1 | P70704 | Channels and Receptors | Other Channels and Receptors | phosphorylated | |
| Slc24a2 | Q14BI1 | Channels and Receptors | Other Channels and Receptors | phosphorylated | |
| Slc6a17 | Q8BJI1 | Channels and Receptors | Other Channels and Receptors | phosphorylated | |
| Chrm5 | Q920H4 | Channels and Receptors | Other ligand receptors | phosphorylated | |
| Gabbr1 | G00000851 | Q9WV18 | Channels and Receptors | Other ligand receptors | dephosphorylated |
| Cacna1b | G00000083 | O55017 | Channels and Receptors | Voltage- and Ligand-gated Ca2+-channel | dephosphorylated |
| Cacna1e | G00000078 | Q61290 | Channels and Receptors | Voltage- and Ligand-gated Ca2+-channel | dephosphorylated |
| Cacnb3 | G00000079 | P54285 | Channels and Receptors | Voltage- and Ligand-gated Ca2+-channel | phosphorylated |
| Cacnb4 | Q8R0S4 | Channels and Receptors | Voltage- and Ligand-gated Ca2+-channel | dephosphorylated | |
| Kcnd2 | Q9Z0V2 | Channels and Receptors | Voltage-gated K+-channel | dephosphorylated | |
| Kcnq2 | G00000090 | Q3UY10 | Channels and Receptors | Voltage-gated K+-channel | phosphorylated |
| Map1a | G00001073 | Q9QYR6 | Cytoskeletal | MAPs | phosphorylated + dephosphorylated |
| Map1b | G00000549 | P14873 | Cytoskeletal | MAPs | phosphorylated + dephosphorylated |
| Map2 | G00000547 | P20357 | Cytoskeletal | MAPs | dephosphorylated |
| Map4 | G00000551 | P27546 | Cytoskeletal | MAPs | phosphorylated |
| Mapt | G00001036 | P10637 | Cytoskeletal | MAPs | phosphorylated + dephosphorylated |
| Ablim1 | G00000574 | Q8K4G5 | Cytoskeletal | Other Cytoskeletal Proteins | dephosphorylated |
| Add1 | G00000569 | Q8K232 | Cytoskeletal | Other Cytoskeletal Proteins | phosphorylated |
| Bsn | G00000957 | O88737 | Cytoskeletal | Other Cytoskeletal Proteins | phosphorylated + dephosphorylated |
| Des | G00000606 | P31001 | Cytoskeletal | Other Cytoskeletal Proteins | dephosphorylated |
| Epb41l1 | G00000593 | Q9Z2H5 | Cytoskeletal | Other Cytoskeletal Proteins | dephosphorylated |
| Epb41l3 | G00001827 | Q9WV92 | Cytoskeletal | Other Cytoskeletal Proteins | dephosphorylated |
| Gas2l1 | Q8JZP9 | Cytoskeletal | Other Cytoskeletal Proteins | dephosphorylated | |
| Nefh | G00000585 | P19246 | Cytoskeletal | Other Cytoskeletal Proteins | phosphorylated + dephosphorylated |
| Nefm | G00000567 | P08553 | Cytoskeletal | Other Cytoskeletal Proteins | dephosphorylated |
| P140 | G00000666 | Q9QWI6 | Cytoskeletal | Other Cytoskeletal Proteins | phosphorylated + dephosphorylated |
| Palm | G00000586 | Q9Z0P4 | Cytoskeletal | Other Cytoskeletal Proteins | dephosphorylated |
| Ppp1r9a | G00000197 | Q7TN74 | Cytoskeletal | Other Cytoskeletal Proteins | dephosphorylated |
| Tnks1bp1 | P58871 | Cytoskeletal | Other Cytoskeletal Proteins | dephosphorylated | |
| Spnb3 | Q68FG2 | Cytoskeletal | Spectrin | phosphorylated | |
| Sptbn1 | G00001795 | Q62261 | Cytoskeletal | Spectrin | dephosphorylated |
| Arhgap23 | Q69ZH9 | G-proteins and Modulators | Modulators | dephosphorylated | |
| Arhgef2 | Q60875 | G-proteins and Modulators | Modulators | phosphorylated | |
| Psd3 | G00001270 | Q2PFD7 | G-proteins and Modulators | Modulators | dephosphorylated |
| Syngap1 | G00000009 | Q9QUH6 | G-proteins and Modulators | Modulators | dephosphorylated |
| Aak1 | G00000881 | Q3UHJ0 | Kinases | Ser/Thr Kinases | dephosphorylated |
| Dclk1 | G00000154 | Q9JLM8 | Kinases | Ser/Thr Kinases | dephosphorylated |
| Mark4 | Q8CIP4 | Kinases | Ser/Thr Kinases | phosphorylated | |
| Prkcg | G00001232 | P63318 | Kinases | Ser/Thr Kinases | dephosphorylated |
| Tnik | G00000880 | P83510 | Kinases | Ser/Thr Kinases | dephosphorylated |
| Abi1 | G00000596 | Q8CBW3 | MAGUKs / Adaptors / Scaffolders | non-PDZ-domain containg scaffolders | dephosphorylated |
| Begain | G00000870 | Q68EF6 | MAGUKs / Adaptors / Scaffolders | non-PDZ-domain containg scaffolders | dephosphorylated |
| Dlgap3 | Q6PFD5 | MAGUKs / Adaptors / Scaffolders | non-PDZ-domain containg scaffolders | phosphorylated | |
| Dmxl2 | G00000057 | Q8BPN8 | MAGUKs / Adaptors / Scaffolders | non-PDZ-domain containg scaffolders | dephosphorylated |
| Itsn1 | G00000777 | Q9Z0R4 | MAGUKs / Adaptors / Scaffolders | non-PDZ-domain containg scaffolders | dephosphorylated |
| Sorbs1 | G00000134 | Q62417 | MAGUKs / Adaptors / Scaffolders | non-PDZ-domain containg scaffolders | dephosphorylated |
| Tirap | Q99JY1 | MAGUKs / Adaptors / Scaffolders | non-PDZ-domain containg scaffolders | dephosphorylated | |
| CortBP1 | G00000572 | Q80Z38 | MAGUKs / Adaptors / Scaffolders | PDZ-domain containing scaffolders | dephosphorylated |
| Dlg4 | G00000004 | Q62108 | MAGUKs / Adaptors / Scaffolders | PDZ-domain containing scaffolders | dephosphorylated |
| Lrrc7 | G00000019 | Q80TE7 | MAGUKs / Adaptors / Scaffolders | PDZ-domain containing scaffolders | dephosphorylated |
| Pclo | G00000864 | Q9QYX7 | MAGUKs / Adaptors / Scaffolders | PDZ-domain containing scaffolders | dephosphorylated |
| Shank3 | G00000121 | Q4ACU6 | MAGUKs / Adaptors / Scaffolders | PDZ-domain containing scaffolders | dephosphorylated |
| C20orf30 | Q8CIB6 | Other transmembrane | Other transmembrane | dephosphorylated | |
| Cldn11 | G00000052 | Q60771 | Other transmembrane | Other transmembrane | dephosphorylated |
| Gja1 | G00000626 | P23242 | Other transmembrane | Other transmembrane | phosphorylated + dephosphorylated |
| Kirrel2 | Q7TSU7 | Other transmembrane | Other transmembrane | phosphorylated | |
| Syt6 | Q9R0N8 | Other transmembrane | Other transmembrane | dephosphorylated | |
| Ppp1r14d | Q7TT52 | Protein Phosphatases | Protein Phosphatases | phosphorylated | |
| Ppp1r9b | G00000191 | Q6R891 | Protein Phosphatases | Protein Phosphatases | dephosphorylated |
| Dpysl2 | G00000441 | O08553 | Signalling molecules and Enzymes | Development | dephosphorylated |
| Gprin1 | G00000334 | Q3UNH4 | Signalling molecules and Enzymes | Development | dephosphorylated |
| Bag3 | Q9JLV1 | Signalling molecules and Enzymes | Heat shock / Chaperones / Chaperonins | dephosphorylated | |
| Nsun7 | Q14AW5 | Signalling molecules and Enzymes | Other enzymes | dephosphorylated | |
| Pde4d | G00000290 | Q01063 | Signalling molecules and Enzymes | Other enzymes | dephosphorylated |
| Pin4 | Q9CWW6 | Signalling molecules and Enzymes | Other enzymes | phosphorylated | |
| Top2b | Q64511 | Signalling molecules and Enzymes | Other enzymes | dephosphorylated | |
| G3bp | G00000253 | Q3UR88 | Signalling molecules and Enzymes | Other signalling molecules | dephosphorylated |
| Kcnip2 | Q3YAB1 | Signalling molecules and Enzymes | Other signalling molecules | phosphorylated | |
| Marcksl1 | P28667 | Signalling molecules and Enzymes | Other signalling molecules | dephosphorylated | |
| Plcb1 | G00000355 | Q9Z1B3 | Signalling molecules and Enzymes | Other signalling molecules | dephosphorylated |
| Ppfia3 | G00001017 | P60469 | Signalling molecules and Enzymes | Other signalling molecules | phosphorylated + dephosphorylated |
| Sipa1l1 | G00000020 | Q8C0T5 | Signalling molecules and Enzymes | Other signalling molecules | dephosphorylated |
| Myo18a | G00000756 | Q9JMH9 | Synaptic Vesicles / Protein Transport | Motor Proteins | dephosphorylated |
| Rims1 | G00001087 | Q99NE5-7 | Synaptic Vesicles / Protein Transport | Synaptic vesicle | dephosphorylated |
| Rpl14 | G00001259 | Q9CR57 | Transcription and Translation | Ribosomal Proteins | phosphorylated |
| Bat2 | Q7TSC1 | Transcription and Translation | Splicing | dephosphorylated | |
| Hnrpa1 | P49312 | Transcription and Translation | Splicing | dephosphorylated | |
| Hnrpk | P61979 | Transcription and Translation | Splicing | phosphorylated | |
| Sfrs10 | P62996 | Transcription and Translation | Splicing | dephosphorylated | |
| Sfrs7 | Q8BL97 | Transcription and Translation | Splicing | dephosphorylated | |
| Srrm1 | Q52KI8 | Transcription and Translation | Splicing | dephosphorylated | |
| Srrm2 | Q8BTI8 | Transcription and Translation | Splicing | dephosphorylated | |
| Tra2a | Q6PFR5 | Transcription and Translation | Splicing | dephosphorylated | |
| Cbx3 | P23198 | Transcription and Translation | Transcription | dephosphorylated | |
| Mll | P55200 | Transcription and Translation | Transcription | dephosphorylated | |
| Pparbp | Q925J9 | Transcription and Translation | Transcription | dephosphorylated | |
| Purb | G00000951 | O35295 | Transcription and Translation | Transcription | dephosphorylated |
| Smarcc2 | Q6PDG5 | Transcription and Translation | Transcription | dephosphorylated | |
| Son | Q9QX47 | Transcription and Translation | Transcription | dephosphorylated | |
| Zkscan17 | Q5SXI5 | Transcription and Translation | Transcription | phosphorylated | |
| Eif3s9 | Q8JZQ9 | Transcription and Translation | Translation | phosphorylated | |
| ... | ... | Uncharacterised / novel | Uncharacterised / novel | dephosphorylated | |
| 2700049A03Rik | Q8C9T9 | Uncharacterised / novel | Uncharacterised / novel | dephosphorylated | |
| 4932438A13Rik | Q05BP8 | Uncharacterised / novel | Uncharacterised / novel | dephosphorylated | |
| 4933411K20Rik | Q9D462 | Uncharacterised / novel | Uncharacterised / novel | dephosphorylated | |
| Bcas1 | G00000821 | Q8C0B4 | Uncharacterised / novel | Uncharacterised / novel | phosphorylated |
| C2orf33 homolog | Q6PCP5 | Uncharacterised / novel | Uncharacterised / novel | dephosphorylated | |
| Col20a1 | Q923P0 | Uncharacterised / novel | Uncharacterised / novel | dephosphorylated | |
| Dnajc6 | G00000282 | Q80TZ3 | Uncharacterised / novel | Uncharacterised / novel | dephosphorylated |
| Efcab5 | Q5NC53 | Uncharacterised / novel | Uncharacterised / novel | phosphorylated | |
| Farp1 | Q3UY58 | Uncharacterised / novel | Uncharacterised / novel | dephosphorylated | |
| Gm1568 | Q3UHB8 | Uncharacterised / novel | Uncharacterised / novel | phosphorylated + dephosphorylated | |
| Gm879 | Q5SSH6 | Uncharacterised / novel | Uncharacterised / novel | dephosphorylated | |
| Hd | G00001247 | P42859 | Uncharacterised / novel | Uncharacterised / novel | dephosphorylated |
| Hdgfrp2 | Q99L92 | Uncharacterised / novel | Uncharacterised / novel | dephosphorylated | |
| Hisppd2a | Q7TSP1 | Uncharacterised / novel | Uncharacterised / novel | dephosphorylated | |
| Kbtbd11 | Q8BNW9 | Uncharacterised / novel | Uncharacterised / novel | dephosphorylated | |
| Kctd16 | Q5DTY9 | Uncharacterised / novel | Uncharacterised / novel | phosphorylated | |
| KIAA0773 homolog | Q3TY60 | Uncharacterised / novel | Uncharacterised / novel | dephosphorylated | |
| mg638 | Q9ER87 | Uncharacterised / novel | Uncharacterised / novel | dephosphorylated | |
| Ndrg2 | Q9CTJ5 | Uncharacterised / novel | Uncharacterised / novel | dephosphorylated | |
| Nucks1 | Q80XU3 | Uncharacterised / novel | Uncharacterised / novel | dephosphorylated | |
| Odf2 | A3KGV1 | Uncharacterised / novel | Uncharacterised / novel | phosphorylated | |
| Sgip1 | G00000827 | Q8VD37 | Uncharacterised / novel | Uncharacterised / novel | dephosphorylated |
| unnamed protein product | BAC39111 | Uncharacterised / novel | Uncharacterised / novel | dephosphorylated | |
| Wapal | Q65Z40 | Uncharacterised / novel | Uncharacterised / novel | dephosphorylated | |
| Wdr20a | Q8R0J5 | Uncharacterised / novel | Uncharacterised / novel | phosphorylated |