G2C::Genetics
TNIK is required for postsynaptic and nuclear signalling pathways and cognitive function
Marcello P. Coba1,2*, Noboru H. Komiyama1,3*, Jess Nithianantharajah1,3*, Maksym V. Kopanitsa1,4, Tim Indersmitten5, Nathan G. Skene1, Ellie J. Tuck1,4, David G. Fricker1,4, Kathryn A. Elsegood1,3, Lianne E. Stanford1, Nurudeen Afinowi1,4, Lisa M. Saksida6, Timothy J. Bussey6, Thomas J. O'Dell7 and Seth G.N. Grant1,3,6
Author email: Seth.Grant@ed.ac.uk * - These authors contributed equally to this work
- Genes to Cognition Programme, The Wellcome Trust Sanger Institute, Genome Campus, Hinxton, Cambridgeshire, CB10 1SA, UK
- Zilkha Neurogenetic Institute, Department of Psychiatry and Behavioral Sciences, University of Southern California, Los Angeles, California, 90089, USA
- Centre for Clinical Brain Sciences and Centre for Neuroregeneration, The University of Edinburgh, Chancellors Building, 47 Little France Crescent, Edinburgh EH16 4SB, UK
- Synome Ltd, Babraham Research Campus, Cambridge, CB22 3AT, UK
- Interdepartmental PhD Program for Neuroscience, UCLA, Los Angeles, California 90095, USA
- Department of Experimental Psychology, University of Cambridge, UK; The MRC and Wellcome Trust Behavioural and Clinical Neuroscience Institute, University of Cambridge, Downing St., Cambridge CB2 3EB, UK
- Department of Physiology, David Geffen School of Medicine at UCLA, Los Angeles, California 90095, USA
Traf2 and NcK interacting Kinase (TNiK) contains serine-threonine kinase and scaffold domains and has been implicated in cell proliferation and glutamate receptor regulation in vitro. Here we report its role in vivo using mice carrying a knockout mutation.
TNiK binds protein complexes in the synapse linking it to the NMDA receptor (NMDAR) via AKAP9. NMDAR and metabotropic receptors bidirectionally regulate TNiK phosphorylation and TNiK was required for AMPA expression and synaptic function. TNiK also organises nuclear complexes and in the absence of TNiK, there was a marked elevation in GSK3Β and phosphorylation levels of its cognate phosphorylation sites on NeuroD1 with alterations in Wnt pathway signalling.
We observed impairments in dentate gyrus neurogenesis in TNiK knockout mice and cognitive testing using the touchscreen apparatus revealed impairments in pattern separation on a test of spatial discrimination. Object-location paired associates learning, which is dependent on glutamatergic signalling was also impaired. Additionally, TNiK knockout mice displayed hyperlocomotor behavior that could be rapidly reversed by GSK3Β inhibitors, indicating the potential for pharmacological rescue of a behavioral phenotype.
These data establish TNiK as a critical regulator of cognitive functions and suggest it may play a regulatory role in diseases impacting on its interacting proteins and complexes.
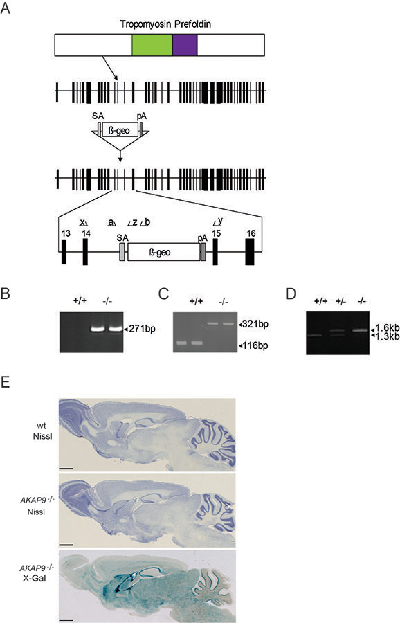
Supplementary Figure 1. Generation of AKAP9 mutant mice
A mouse embryonic stem (ES) cell line (XP0050, strain 129/Ola) with an insertional mutation in Akap9 was obtained from Sanger Institute Gene Trap Resource (SIGTR - sanger.ac.uk/PostGenomics/genetrap/). The insertional mutation in XP0050 by the gene-trapping vector, pGT0lxr, was designed to create an in-frame fusion between the 5' exons of the trapped gene and a reporter, Β-geo (a fusion of Β-galactosidase and neomycin phosphotransferase II) occurred in intron 14-15 (Transcript: Akap9-001 (ENSMUST00000044492) Ensembl release 56). Thus, the gene-trapped locus is predicted to yield a fusion transcript containing exons 1-14 of Akap9 and Β-geo. Integration of the gene-trapping vector was confirmed by RT-PCR. A 217bp product was amplified from the gene trap clone cDNA using primer x (GAAGCTGTCTAAGAGAGTGTG) that hybridises the sequence encoded by the part of exon 14 with reverse Primer z (GATCCTCTAGAGTCCAGATCTG) within the Β-geo cassette. This gene trap ES cell clone was injected into C57/BL6 blastocysts to create chimeric mice, which were bred with 129S5 mice to generate heterozygous (+/–) Akap9 mutant mice. Those F1 heterozygous mice had been backcrossed with 129S5 mice for 1-2 times before being used for intercrossing.
A. Gene trap vector for the generation of AKAP9 knockout mice. Linear structure of AKAP9 protein domain (top diagram). AKAP9 is a 48 exon protein. The gene-trapping vector, pGT0lxr, is inserted between exon14 and 15. Primers used for genotyping (see supplementary methods) are shown (bottom diagram). SA, splice acceptor; pA, polyadenylation signal; Β-geo, fusion of Β-galactosidase and neomycin phosphotransferase II.
B. Electrophoresis gel image showing confirmation by RTPCR that trap is correctly inserted.
C. RT-PCR genotyping. Homozygote (-/-) product is 205bp larger than wt (+/+) product.
D. PCR genotyping of targeted AKAP9-/- mice using a common forward primer, a, and reverse primers b and y to amplify the wt and mutant alleles respectively.
Supplementary Table 1. Hippocampal mRNA expression profile of TNiK-/- mice
Shown are the 200 most significant genes from the mRNA expression profiling, the whole genome array results are downloadable.
Download whole genome microarray results
| Gene Symbol | Gene Description | Ensembl Gene ID | Entrez Gene ID | Average Expression | log Fold Change | P Value | Adjusted P Value | t | Genomic Location | Probe Location | RefSeq transcripts | Probe id |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Spata13 | Spermatogenesis associated 13 | ENSMUSG00000021990 | 9.20 | -0.357 | 6.44E-09 | 1.49E-04 | -7.17 | chr14:61383157:61383206:+ | 3pUTR | XM_923227 XM_901902 | ILMN_2751037 | |
| Cort | Cortistatin | ENSMUSG00000028971 | 258453 | 8.40 | 0.378 | 3.32E-08 | 3.84E-04 | 6.69 | chr4:148499369:148499418:- | CDS | NM_007745 | ILMN_1226607 |
| BC050868 | Mus musculus adult male cerebellum cDNA, RIKEN full-length enriched library, clone:1500031A17 product:TRAF2 AND NCK INTERACTING KINASE, SPLICE VARIANT 8 homolog [Homo sapiens], full insert sequence. | ENSMUSG00000027692 | 10.07 | -0.403 | 6.85E-08 | 5.28E-04 | -6.47 | chr3:28569142:28569191:+ | 3pUTR | XM_001473621 XM_001473602 XM_001474909 XM_001474897 | ILMN_2524251 | |
| C4b | Complement component 4B (Childo blood group) | ENSMUSG00000073418 | 212285 | 8.68 | -0.335 | 3.40E-06 | 1.66E-02 | -5.32 | chr17:34866134:34866183:- | CDS | NM_009780 NM_011413 XM_973028 XM_972987 XM_973068 | ILMN_1215092 |
| Scara3 | Scavenger receptor class A, member 3 | ENSMUSG00000034463 | 268996 | 10.26 | -0.283 | 5.07E-06 | 1.66E-02 | -5.20 | chr14:66538293:66538342:- | 3pUTR | NM_172604 | ILMN_2706268 |
| Slc30a3 | Solute carrier family 30 (zinc transporter), member 3 | ENSMUSG00000029151 | 22784 | 12.22 | -0.283 | 5.20E-06 | 1.66E-02 | -5.19 | chr5:31388548:31388597:- | 3pUTR | ILMN_2521965 | |
| C4b | Complement component 4B (Childo blood group) | ENSMUSG00000015451 | 78829 | 8.53 | -0.288 | 5.82E-06 | 1.66E-02 | -5.15 | chr17:34865420:34865469:- | CDS | NM_009780 NM_011413 XM_973028 XM_972987 XM_973068 | ILMN_3049559 |
| Tcirg1 | T-cell, immune regulator 1, ATPase, H+ transporting, lysosomal V0 protein A3 | ENSMUSG00000001750 | 258203 | 7.57 | -0.103 | 6.47E-06 | 1.66E-02 | -5.12 | chr19:3903583:3903632:- | CDS | NM_016921 | ILMN_2643879 |
| Hn1 | Hematological and neurological expressed sequence 1 | ENSMUSG00000020737 | 69202 | 12.59 | -0.196 | 6.80E-06 | 1.66E-02 | -5.11 | chr11:115358689:115358738:- | 3pUTR | NM_008258 | ILMN_2914744 |
| Cox6a2 | Cytochrome c oxidase, subunit VI a, polypeptide 2 | ENSMUSG00000030785 | 258289 | 8.53 | 0.336 | 7.19E-06 | 1.66E-02 | 5.09 | chr7:135349162:135349211:- | 3pUTR | NM_009943 | ILMN_2629581 |
| 9130213B05Rik | RIKEN cDNA 9130213B05 gene | ENSMUSG00000034981 | 231440 | 11.20 | 0.437 | 1.20E-05 | 2.23E-02 | 4.93 | chr5:92052926:92052975:+ | 3pUTR | NM_145562 | ILMN_2756704 |
| Ebf3 | Early B-cell factor 3 | ENSMUSG00000010476 | 7.95 | 0.127 | 1.22E-05 | 2.23E-02 | 4.93 | chr7:144385579:144385628:- | 3pUTR | NM_001113414 NM_001113415 NM_010096 | ILMN_2422615 | |
| Rhag | Rhesus blood group-associated A glycoprotein | ENSMUSG00000023926 | 16428 | 7.55 | -0.084 | 1.25E-05 | 2.23E-02 | -4.92 | chr17:40971620:40971669:+ | CDS | NM_011269 | ILMN_2745515 |
| Rabggtb | RAB geranylgeranyl transferase, b subunit | ENSMUSG00000038975 | 338360 | 7.74 | -0.146 | 1.64E-05 | 2.70E-02 | -4.84 | chr3:153573046:153573095:- | 3pUTR | ILMN_2703263 | |
| Myo5b | Myosin VB | ENSMUSG00000025885 | 17919 | 9.94 | -0.423 | 2.07E-05 | 3.19E-02 | -4.77 | chr18:74930039:74930088:+ | CDS | NM_201600 | ILMN_2539489 |
| A830018L16Rik | RIKEN cDNA A830018L16 gene | 69035 | 8.72 | -0.210 | 2.23E-05 | 3.23E-02 | -4.75 | chr1:11588290:11588339:+ | 3pUTR | ILMN_1213044 | ||
| Nat8l | N-acetyltransferase 8-like | ENSMUSG00000048142 | 56208 | 9.72 | 0.284 | 2.59E-05 | 3.53E-02 | 4.70 | chr5:34345517:34345566:+ | 3pUTR | NM_001001985 | ILMN_2594039 |
| Grasp | GRP1 (general receptor for phosphoinositides 1)-associated scaffold protein | ENSMUSG00000000531 | 67692 | 11.35 | -0.216 | 2.84E-05 | 3.64E-02 | -4.67 | chr15:101063106:101063155:+ | 3pUTR | NM_019518 | ILMN_2596396 |
| Usp52 | Ubiquitin specific peptidase 52 | ENSMUSG00000005682 | 74559 | 8.57 | -0.254 | 3.42E-05 | 3.66E-02 | -4.61 | chr10:127758185:127758234:+ | 3pUTR | NM_133992 | ILMN_2790392 |
| Bat2d | BAT2 domain containing 1 | ENSMUSG00000040225 | 71983 | 7.82 | 0.305 | 3.67E-05 | 3.66E-02 | 4.59 | chr1:164640509:164640558:- | CDS | NM_001081290 | ILMN_1239599 |
| Lum | Lumican | ENSMUSG00000036446 | 8.24 | -0.279 | 3.72E-05 | 3.66E-02 | -4.59 | chr10:97034583:97034632:+ | CDS | NM_008524 | ILMN_3001540 | |
| Tle1 | Transducin-like enhancer of split 1, homolog of Drosophila E(spl) | ENSMUSG00000008305 | 240667 | 8.62 | -0.275 | 4.25E-05 | 3.66E-02 | -4.55 | chr4:71819008:71819057:- | 3pUTR | ILMN_1245089 | |
| Ifnz | Interferon zeta | ENSMUSG00000073810 | 74753 | 7.59 | -0.089 | 4.58E-05 | 3.66E-02 | -4.52 | chr4:88428166:88428215:+ | 5pUTR | NM_197889 | ILMN_3021483 |
| B4galnt4 | Beta-1,4-N-acetyl-galactosaminyl transferase 4 | 330671 | 8.51 | -0.178 | 4.67E-05 | 3.66E-02 | -4.52 | chr7:148254691:148254740:+ | CDS | ILMN_2540866 | ||
| Zzz3 | Zinc finger, ZZ domain containing 3 | ENSMUSG00000039068 | 15481 | 7.61 | 0.093 | 4.68E-05 | 3.66E-02 | 4.52 | chr3:152085636:152085685:+ | 5pUTR | XR_001912 XR_032051 NM_198416 NM_001080755 | ILMN_2424408 |
| Scara5 | Scavenger receptor class A, member 5 (putative) | ENSMUSG00000022032 | 7.55 | -0.088 | 4.83E-05 | 3.66E-02 | -4.51 | chr14:66383255:66383304:+ | 3pUTR | NM_028903 | ILMN_3008068 | |
| Htr1f | 5-hydroxytryptamine (serotonin) receptor 1F | ENSMUSG00000050783 | 227682 | 7.86 | 0.182 | 4.96E-05 | 3.66E-02 | 4.50 | chr16:64924829:64924878:- | 3pUTR | NM_008310 | ILMN_2933261 |
| Eltd1 | EGF, latrophilin seven transmembrane domain containing 1 | ENSMUSG00000039167 | 140577 | 7.96 | 0.162 | 4.97E-05 | 3.66E-02 | 4.50 | chr3:151207706:151207755:+ | 3pUTR | NM_133222 | ILMN_1224540 |
| Stx4a | Syntaxin 4A (placental) | ENSMUSG00000030805 | 228880 | 8.50 | 0.158 | 5.49E-05 | 3.66E-02 | 4.47 | chr7:134992177:134992181:+,chr7:134992265:134992309:+ | 3pUTR | NM_009294 | ILMN_2822053 |
| 2810006K23Rik | RIKEN cDNA 2810006K23 gene | ENSMUSG00000047635 | 12661 | 7.58 | 0.073 | 5.53E-05 | 3.66E-02 | 4.46 | chr5:124790696:124790745:+ | CDS | ILMN_2546984 | |
| Mtap1a | Microtubule-associated protein 1 A | ENSMUSG00000027254 | 74143 | 8.14 | 0.315 | 5.66E-05 | 3.66E-02 | 4.46 | chr2:121126148:121126197:+ | CDS | NM_032393 | ILMN_2646070 |
| EG434171 | Predicted gene, EG434171 | ENSMUSG00000060565 | 76486 | 7.69 | -0.115 | 6.06E-05 | 3.66E-02 | -4.44 | chr7:39303890:39303939:- | CDS | NM_001013810 | ILMN_2879144 |
| Ptprb | Protein tyrosine phosphatase, receptor type, B | ENSMUSG00000020154 | 27357 | 8.10 | 0.388 | 6.19E-05 | 3.66E-02 | 4.43 | chr10:115820928:115820977:+ | CDS | NM_029928 | ILMN_2591731 |
| Cort | Cortistatin | ENSMUSG00000028971 | 238323 | 8.70 | 0.349 | 6.39E-05 | 3.66E-02 | 4.42 | chr4:148499316:148499365:- | 3pUTR | NM_007745 | ILMN_2955725 |
| 2310007H09Rik | RIKEN cDNA 2310007H09 gene | ENSMUSG00000030946 | 258293 | 8.60 | -0.219 | 6.40E-05 | 3.66E-02 | -4.42 | chr7:139897331:139897380:+ | CDS | NM_029609 | ILMN_2600720 |
| AK021368 | Mus musculus 0 day neonate eyeball cDNA, RIKEN full-length enriched library, clone:E130102H24 product:hypothetical protein, full insert sequence. | 18105 | 8.45 | -0.313 | 6.54E-05 | 3.66E-02 | -4.41 | chr4:101019192:101019241:- | 3pUTR | XM_001472210 XM_992447 | ILMN_2647694 | |
| Ddrgk1 | DDRGK domain containing 1 | ENSMUSG00000068290 | 11.57 | -0.207 | 6.54E-05 | 3.66E-02 | -4.41 | chr2:130479850:130479899:- | CDS | NM_029832 | ILMN_2874128 | |
| Parp2 | Poly (ADP-ribose) polymerase family, member 2 | ENSMUSG00000036023 | 11546 | 9.96 | -0.154 | 6.61E-05 | 3.66E-02 | -4.41 | chr14:51440785:51440834:+ | CDS | NM_009632 | ILMN_1226731 |
| Gm1040 | Gene model 1040, (NCBI) | ENSMUSG00000001569 | 433864 | 7.62 | -0.095 | 6.80E-05 | 3.66E-02 | -4.40 | chr5:29762076:29762125:+ | CDS | NM_001033457 | ILMN_2537214 |
| Bcor | BCL6 interacting corepressor | ENSMUSG00000040363 | 68720 | 8.08 | -0.177 | 6.89E-05 | 3.66E-02 | -4.40 | chrX:11614985:11615034:- | CDS | NM_175045 NM_029510 NM_175044 NM_175046 | ILMN_1228091 |
| Ankrd11 | Ankyrin repeat domain 11 | ENSMUSG00000035569 | 8.16 | 0.386 | 6.89E-05 | 3.66E-02 | 4.40 | chr8:125417578:125417627:- | CDS | NM_001081379 | ILMN_2565729 | |
| Shmt2 | Serine hydroxymethyltransferase 2 (mitochondrial) | 8.07 | 0.118 | 7.03E-05 | 3.66E-02 | 4.39 | chr10:126954204:126954253:- | 3pUTR | NM_028230 | ILMN_1224942 | ||
| Bat2d | BAT2 domain containing 1 | ENSMUSG00000040225 | 7.99 | 0.432 | 7.12E-05 | 3.66E-02 | 4.39 | chr1:164640508:164640557:- | CDS | NM_001081290 | ILMN_1241229 | |
| Irx3 | Iroquois related homeobox 3 (Drosophila) | ENSMUSG00000031734 | 74895 | 7.61 | -0.078 | 7.24E-05 | 3.66E-02 | -4.38 | chr8:94322817:94322866:- | CDS | NM_008393 | ILMN_2776909 |
| Psen1 | Presenilin 1 | ENSMUSG00000019969 | 58234 | 8.65 | -0.308 | 7.48E-05 | 3.66E-02 | -4.37 | chr12:85074933:85074982:+ | CDS | NM_008943 | ILMN_1214811 |
| Slc7a7 | Solute carrier family 7 (cationic amino acid transporter, y+ system), member 7 | ENSMUSG00000000958 | 218103 | 7.66 | -0.101 | 7.61E-05 | 3.66E-02 | -4.37 | chr14:54992648:54992697:- | CDS | NM_011405 | ILMN_2723826 |
| Bccip | BRCA2 and CDKN1A interacting protein | ENSMUSG00000030983 | 67606 | 10.59 | -0.253 | 7.69E-05 | 3.66E-02 | -4.36 | chr7:140912451:140912500:+ | CDS | NM_025392 | ILMN_2759920 |
| C330006K01Rik | RIKEN cDNA C330006K01 gene | ENSMUSG00000039623 | 8.40 | 0.108 | 8.18E-05 | 3.66E-02 | 4.34 | chr5:142953681:142953730:+ | 3pUTR | XM_001480714 XM_001480711 XM_001480708 NM_172725 | ILMN_2752206 | |
| Nras | Neuroblastoma ras oncogene | 14158 | 10.94 | -0.315 | 8.41E-05 | 3.66E-02 | -4.33 | chr3:102871584:102871633:+ | 3pUTR | NM_010937 | ILMN_2837543 | |
| Ppp2r4 | Protein phosphatase 2A, regulatory subunit B (PR 53) | ENSMUSG00000039515 | 216881 | 8.28 | 0.209 | 8.41E-05 | 3.66E-02 | 4.33 | chr2:30303115:30303164:+ | 3pUTR | NM_138748 | ILMN_2755399 |
| Acpl2 | Acid phosphatase-like 2 | ENSMUSG00000043587 | 9.59 | -0.285 | 8.59E-05 | 3.66E-02 | -4.33 | chr9:96723879:96723928:- | 3pUTR | NM_153420 | ILMN_1254634 | |
| 2810474O19Rik | RIKEN cDNA 2810474O19 gene | ENSMUSG00000032712 | 67246 | 7.81 | 0.217 | 8.68E-05 | 3.66E-02 | 4.32 | chr6:149274925:149274974:+ | CDS | NM_026054 | ILMN_2596077 |
| Sqstm1 | Sequestosome 1 | ENSMUSG00000015837 | 18048 | 11.70 | -0.350 | 8.85E-05 | 3.66E-02 | -4.32 | chr11:50014169:50014218:- | 3pUTR | NM_011018 | ILMN_2710705 |
| Fhl1 | Four and a half LIM domains 1 | ENSMUSG00000023092 | 14615 | 11.02 | -0.319 | 8.89E-05 | 3.66E-02 | -4.32 | chrX:54046293:54046342:+ | 3pUTR | NM_010211 NM_001077361 NM_001077362 | ILMN_2713285 |
| Dennd5a | DENN/MADD domain containing 5A | ENSMUSG00000035901 | 15951 | 11.05 | -0.224 | 9.32E-05 | 3.66E-02 | -4.30 | chr7:117037678:117037727:- | 3pUTR | NM_021494 XM_001477127 | ILMN_3107607 |
| Lcat | Lecithin cholesterol acyltransferase | ENSMUSG00000035237 | 16816 | 8.38 | 0.164 | 9.36E-05 | 3.66E-02 | 4.30 | chr8:108463676:108463725:- | CDS | NM_008490 | ILMN_2798400 |
| Hmgn2 | High mobility group nucleosomal binding domain 2 | ENSMUSG00000003038 | 10.90 | -0.338 | 9.43E-05 | 3.66E-02 | -4.30 | chr4:133521188:133521236:- | 3pUTR | XM_001478555 | ILMN_1238360 | |
| Hmbs | Hydroxymethylbilane synthase | ENSMUSG00000073552 | 70120 | 8.72 | -0.222 | 9.44E-05 | 3.66E-02 | -4.30 | chr9:44145454:44145503:- | CDS | XM_915480 NM_013551 XM_888318 NM_001110251 | ILMN_2657124 |
| Lmo3 | LIM domain only 3 | ENSMUSG00000030226 | 13090 | 10.91 | -0.273 | 9.54E-05 | 3.66E-02 | -4.29 | chr6:138313349:138313398:- | 3pUTR | NM_207222 | ILMN_3062075 |
| Flt1 | FMS-like tyrosine kinase 1 | ENSMUSG00000029648 | 14254 | 9.72 | 0.444 | 9.65E-05 | 3.66E-02 | 4.29 | chr5:148373789:148373838:- | 3pUTR | NM_010228 | ILMN_1227926 |
| Zfp821 | Zinc finger protein 821 | ENSMUSG00000031728 | 319801 | 8.36 | -0.300 | 9.66E-05 | 3.66E-02 | -4.29 | chr8:112246886:112246935:+ | CDS | NM_029468 | ILMN_2693387 |
| Pdlim3 | PDZ and LIM domain 3 | 23834 | 7.79 | 0.118 | 9.92E-05 | 3.70E-02 | 4.28 | chr8:47004761:47004810:+ | 3pUTR | NM_016798 | ILMN_2885990 | |
| Src | Rous sarcoma oncogene | ENSMUSG00000027646 | 245308 | 7.71 | -0.094 | 0.000101784 | 3.74E-02 | -4.27 | chr2:157295241:157295257:+,chr2:157295489:157295521:+ | CDS | NM_001025395 NM_009271 | ILMN_2677529 |
| Ankra2 | Ankyrin repeat, family A (RFXANK-like), 2 | ENSMUSG00000021661 | 26428 | 8.57 | -0.248 | 0.000104098 | 3.76E-02 | -4.27 | chr13:99043230:99043279:+ | 3pUTR | ILMN_1243090 | |
| Hmgn2 | High mobility group nucleosomal binding domain 2 | ENSMUSG00000003038 | 10.94 | -0.426 | 0.000112371 | 3.86E-02 | -4.24 | chr4:133521180:133521229:- | 3pUTR | XM_001473104 XM_001477164 XM_001476615 XM_891577 NM_016957 XM_001478560 XM_001479392 | ILMN_1237216 | |
| Prss23 | Protease, serine, 23 | 76453 | 7.60 | 0.080 | 0.000114291 | 3.86E-02 | 4.24 | chr7:96656349:96656398:- | 3pUTR | NM_029614 | ILMN_1246127 | |
| Klhl6 | Kelch-like 6 (Drosophila) | ENSMUSG00000043008 | 382406 | 7.58 | 0.079 | 0.000114985 | 3.86E-02 | 4.23 | chr16:19953586:19953635:- | CDS | NM_183390 | ILMN_1260233 |
| Slco1c1 | Solute carrier organic anion transporter family, member 1c1 | ENSMUSG00000030235 | 12033 | 9.58 | 0.378 | 0.000116713 | 3.86E-02 | 4.23 | chr6:141517967:141518016:+ | CDS | NM_021471 | ILMN_1242173 |
| Cog5 | 6 days neonate head cDNA, RIKEN full-length enriched library, clone:5430405C01 product:PUTATIVE 13 S GOLGI TRANSPORT COMPLEX 90KD SUBUNIT BRAIN-SPECIFIC ISOFORM homolog (Fragment). | ENSMUSG00000035933 | 226352 | 8.63 | -0.196 | 0.000119492 | 3.86E-02 | -4.22 | chr12:32604642:32604642:+,chr12:32605419:32605467:+ | CDS | ILMN_2523980 | |
| Krt14 | Keratin 14 | ENSMUSG00000045545 | 259027 | 7.57 | 0.075 | 0.000120037 | 3.86E-02 | 4.22 | chr11:100064523:100064572:- | 3pUTR | NM_016958 | ILMN_2722616 |
| Fbxl6 | F-box and leucine-rich repeat protein 6 | ENSMUSG00000022559 | 76937 | 8.80 | -0.212 | 0.000120243 | 3.86E-02 | -4.22 | chr15:76367007:76367056:- | CDS | NM_013909 | ILMN_1216206 |
| Sqle | Squalene epoxidase | ENSMUSG00000022351 | 66988 | 10.56 | -0.358 | 0.000123862 | 3.86E-02 | -4.21 | chr15:59161327:59161376:+ | CDS | NM_009270 | ILMN_2737163 |
| Inhba | Inhibin beta-A | ENSMUSG00000041324 | 11610 | 8.11 | 0.208 | 0.000124035 | 3.86E-02 | 4.21 | chr13:16118809:16118858:+ | CDS | NM_008380 | ILMN_2868220 |
| LOC380728 | similar to potassium voltage-gated channel, | ENSMUSG00000035355 | 380728 | 7.59 | -0.097 | 0.000127193 | 3.86E-02 | -4.20 | chr11:100605448:100605497:- | CDS | NM_001081194 | ILMN_2534442 |
| Pkd1l2 | Polycystic kidney disease 1 like 2 | 76645 | 7.60 | -0.085 | 0.000127442 | 3.86E-02 | -4.20 | chr8:119547574:119547623:- | 3pUTR | ILMN_2550168 | ||
| Esr1 | Estrogen receptor 1 (alpha) | ENSMUSG00000019768 | 74090 | 7.61 | 0.087 | 0.000127734 | 3.86E-02 | 4.20 | chr10:5342933:5342982:- | 3pUTR | NM_007956 | ILMN_2726412 |
| Cpd | Carboxypeptidase D | ENSMUSG00000020841 | 71083 | 7.67 | 0.122 | 0.000128508 | 3.86E-02 | 4.20 | chr11:76595547:76595596:- | 3pUTR | NM_007754 XM_001476801 | ILMN_2727598 |
| Lypd6 | LY6/PLAUR domain containing 6 | ENSMUSG00000050447 | 12661 | 8.42 | 0.230 | 0.00013045 | 3.87E-02 | 4.19 | chr2:50047963:50048012:+ | 3pUTR | NM_177139 | ILMN_2729745 |
| Pfdn5 | Prefoldin 5 | ENSMUSG00000001289 | 271209 | 12.67 | -0.208 | 0.000134027 | 3.90E-02 | -4.19 | chr15:102158963:102159012:+ | CDS | NM_027044 | ILMN_2671747 |
| 9130011J15Rik | RIKEN cDNA 9130011J15 gene | 100038859 | 10.60 | -0.281 | 0.000134826 | 3.90E-02 | -4.18 | chr8:75090014:75090063:- | 3pUTR | NM_172396 | ILMN_2593484 | |
| Atp6v1b1 | ATPase, H+ transporting, lysosomal V1 subunit B1 | ENSMUSG00000006269 | 104111 | 7.65 | -0.074 | 0.000142094 | 4.04E-02 | -4.17 | chr6:83708597:83708646:+ | 3pUTR | NM_134157 | ILMN_2879910 |
| Epb4.1l5 | Erythrocyte protein band 4.1-like 5 | 226352 | 7.53 | -0.068 | 0.000143127 | 4.04E-02 | -4.17 | chr1:121491758:121491807:- | 3pUTR | NM_001113416 | ILMN_2586966 | |
| Cln3 | Ceroid lipofuscinosis, neuronal 3, juvenile (Batten, Spielmeyer-Vogt disease) | ENSMUSG00000030720 | 245684 | 8.85 | -0.207 | 0.000150228 | 4.12E-02 | -4.15 | chr7:133715628:133715677:- | 3pUTR | XM_001479838 NM_009907 | ILMN_1214952 |
| Heph | Hephaestin | ENSMUSG00000031209 | 224824 | 7.95 | -0.179 | 0.000151892 | 4.12E-02 | -4.15 | chrX:93769002:93769051:+ | CDS | NM_010417 | ILMN_2734252 |
| C4b | Complement component 4B (Childo blood group) | ENSMUSG00000073418 | 67978 | 7.58 | -0.076 | 0.000152264 | 4.12E-02 | -4.15 | chr17:34871083:34871132:- | CDS | NM_009780 XM_973028 XM_972987 XM_973068 XM_001475152 | ILMN_2632567 |
| Vmn2r111 | Vomeronasal 2, receptor 111 | ENSMUSG00000051877 | 210876 | 7.55 | -0.073 | 0.000155285 | 4.12E-02 | -4.14 | chr17:22755892:22755941:+ | CDS | NM_001104575 | ILMN_2530517 |
| Sec22a | SEC22 vesicle trafficking protein homologe A (S. cerevisiae) | ENSMUSG00000034473 | 330177 | 7.60 | 0.054 | 0.000158208 | 4.12E-02 | 4.13 | chr16:35311501:35311550:- | 3pUTR | NM_133704 | ILMN_2742128 |
| Arid4a | AT rich interactive domain 4A (RBP1-like) | ENSMUSG00000048118 | 8.09 | 0.343 | 0.000158314 | 4.12E-02 | 4.13 | chr12:72164095:72164144:+ | CDS | NM_001081195 | ILMN_1239608 | |
| Rps6kl1 | Ribosomal protein S6 kinase-like 1 | ENSMUSG00000019235 | 10.68 | -0.254 | 0.00015841 | 4.12E-02 | -4.13 | chr12:86476728:86476777:- | 3pUTR | NM_146244 | ILMN_2649333 | |
| Lyrm2 | LYR motif containing 2 | 74626 | 7.54 | 0.069 | 0.000169106 | 4.27E-02 | 4.11 | chr4:32888978:32889027:+ | 3pUTR | NM_175364 | ILMN_2732585 | |
| Yif1b | Yip1 interacting factor homolog B (S. cerevisiae) | ENSMUSG00000030588 | 100039496 | 9.72 | -0.266 | 0.000170365 | 4.27E-02 | -4.11 | chr7:30030935:30030984:+ | CDS | NM_001110201 NM_029887 | ILMN_2607608 |
| Timm23 | Translocase of inner mitochondrial membrane 23 homolog (yeast) | ENSMUSG00000069622 | 16842 | 7.60 | 0.092 | 0.000170934 | 4.27E-02 | 4.11 | chr14:32993356:32993405:- | 3pUTR | NM_016897 | ILMN_2646105 |
| Fis1 | Fission 1 (mitochondrial outer membrane) homolog (yeast) | ENSMUSG00000019054 | 68316 | 11.54 | -0.221 | 0.000171757 | 4.27E-02 | -4.11 | chr5:137440995:137441039:+,chr5:137441435:137441439:+ | CDS | NM_025562 | ILMN_2939295 |
| Spn | Sialophorin | ENSMUSG00000051457 | 384572 | 7.67 | -0.091 | 0.000174331 | 4.29E-02 | -4.10 | chr7:134279721:134279770:- | CDS | NM_001037810 NM_009259 | ILMN_3163543 |
| Col25a1 | Collagen, type XXV, alpha 1 | ENSMUSG00000058897 | 77018 | 7.65 | -0.128 | 0.000179504 | 4.33E-02 | -4.09 | chr3:130298747:130298796:+ | 3pUTR | NM_198711 | ILMN_2643186 |
| N4bp2l1 | NEDD4 binding protein 2-like 1 | ENSMUSG00000041132 | 228769 | 10.11 | -0.208 | 0.000179622 | 4.33E-02 | -4.09 | chr5:151374448:151374497:- | 3pUTR | NM_133898 | ILMN_1224250 |
| Lrrc48 | Leucine rich repeat containing 48 | ENSMUSG00000056598 | 74665 | 7.75 | 0.119 | 0.000182667 | 4.33E-02 | 4.09 | chr11:60207649:60207698:+ | CDS | NM_029044 | ILMN_2765593 |
| Adck1 | AarF domain containing kinase 1 | ENSMUSG00000021044 | 30948 | 9.00 | -0.283 | 0.000187562 | 4.33E-02 | -4.08 | chr12:89699685:89699734:+ | 3pUTR | NM_028105 | ILMN_2656440 |
| Spink10 | Serine peptidase inhibitor, Kazal type 10 | ENSMUSG00000044176 | 102442 | 7.80 | 0.157 | 0.000188588 | 4.33E-02 | 4.08 | chr18:62819573:62819622:+ | CDS | NM_177829 | ILMN_2694785 |
| Pdik1l | PDLIM1 interacting kinase 1 like | ENSMUSG00000050890 | 258874 | 7.98 | 0.196 | 0.00018862 | 4.33E-02 | 4.08 | chr4:133831253:133831302:- | 3pUTR | NM_146156 | ILMN_1220174 |
| Fgd6 | FYVE, RhoGEF and PH domain containing 6 | ENSMUSG00000020021 | 66291 | 7.78 | 0.111 | 0.000189312 | 4.33E-02 | 4.08 | chr10:93604599:93604648:+ | 3pUTR | NM_053072 | ILMN_2894961 |
| Nefh | Neurofilament, heavy polypeptide | ENSMUSG00000020396 | 73234 | 9.11 | 0.394 | 0.00019283 | 4.37E-02 | 4.07 | chr11:4839040:4839089:- | 3pUTR | XM_001474024 NM_010904 | ILMN_1237729 |
| Zfp91 | Zinc finger protein 91 | ENSMUSG00000079415 | 69511 | 8.18 | 0.322 | 0.000195072 | 4.38E-02 | 4.07 | chr19:12844870:12844919:- | CDS | NM_001039718 NM_053009 | ILMN_2757889 |
| Pbx1 | Pre B-cell leukemia transcription factor 1 | 258665 | 7.96 | -0.297 | 0.000200103 | 4.44E-02 | -4.06 | chr1:170088153:170088202:- | 3pUTR | XM_001003209 NM_183355 | ILMN_2587711 | |
| Zic3 | Zinc finger protein of the cerebellum 3 | ENSMUSG00000067860 | 7.88 | 0.152 | 0.000202924 | 4.44E-02 | 4.05 | chrX:55289010:55289059:+ | 3pUTR | NM_009575 | ILMN_2440242 | |
| Aars2 | Alanyl-tRNA synthetase 2, mitochondrial (putative) | ENSMUSG00000023938 | 11569 | 7.86 | 0.115 | 0.000203523 | 4.44E-02 | 4.05 | chr17:45657535:45657584:+ | 3pUTR | NM_198608 | ILMN_2876302 |
| Zfp560 | Zinc finger protein 560 | ENSMUSG00000045519 | 22634 | 8.33 | -0.288 | 0.000208354 | 4.48E-02 | -4.04 | chr9:20189489:20189538:- | 5pUTR | NM_001004190 XM_001476734* XM_001474893* | ILMN_1258594 |
| Zfp330 | Zinc finger protein 330 | ENSMUSG00000031711 | 30932 | 8.88 | -0.218 | 0.000210231 | 4.48E-02 | -4.04 | chr8:85288299:85288348:- | CDS | NM_145600 | ILMN_2440789 |
| Fasn | Fatty acid synthase | ENSMUSG00000025153 | 7.63 | -0.104 | 0.000211132 | 4.48E-02 | -4.04 | chr11:120667910:120667959:- | 3pUTR | NM_007988 | ILMN_2874104 | |
| Ddrgk1 | DDRGK domain containing 1 | ENSMUSG00000068290 | 258286 | 11.78 | -0.218 | 0.000219206 | 4.61E-02 | -4.03 | chr2:130479828:130479877:- | 3pUTR | NM_029832 | ILMN_2615854 |
| Elfn1 | Leucine rich repeat and fibronectin type III, extracellular 1 | ENSMUSG00000048988 | 12564 | 8.31 | 0.139 | 0.000225394 | 4.66E-02 | 4.02 | chr5:140450360:140450409:+ | 3pUTR | NM_175522 | ILMN_2606433 |
| Dtx1 | Deltex 1 homolog (Drosophila) | ENSMUSG00000029603 | 8.94 | 0.345 | 0.000225648 | 4.66E-02 | 4.02 | chr5:121130444:121130493:- | 3pUTR | NM_008052 | ILMN_2641793 | |
| Slc35e1 | Solute carrier family 35, member E1 | ENSMUSG00000019731 | 70650 | 9.00 | -0.288 | 0.000229151 | 4.68E-02 | -4.01 | chr8:75005695:75005744:- | 3pUTR | NM_177766 | ILMN_2752618 |
| Pde8b | Phosphodiesterase 8B | ENSMUSG00000021684 | 20044 | 7.69 | -0.106 | 0.000232306 | 4.68E-02 | -4.01 | chr13:95855134:95855183:- | CDS | ILMN_2428320 | |
| Qars | Glutaminyl-tRNA synthetase | ENSMUSG00000032604 | 75424 | 8.89 | -0.200 | 0.000234335 | 4.68E-02 | -4.01 | chr9:108416882:108416931:+ | CDS | NM_133794 | ILMN_2989263 |
| Lrrc4c | Leucine rich repeat containing 4C | ENSMUSG00000050587 | 16182 | 7.59 | 0.081 | 0.000234559 | 4.68E-02 | 4.01 | chr2:97471041:97471090:+ | CDS | NM_178725 | ILMN_1247141 |
| 1700018C11Rik | RIKEN cDNA 1700018C11 gene | ENSMUSG00000058935 | 75524 | 7.54 | -0.078 | 0.000238298 | 4.71E-02 | -4.00 | chr4:63268306:63268355:- | CDS | NM_029324 | ILMN_2649579 |
| Fam3b | Family with sequence similarity 3, member B | ENSMUSG00000022938 | 330319 | 7.54 | 0.060 | 0.000249591 | 4.73E-02 | 3.99 | chr16:97697342:97697352:-,chr16:97696949:97696987:- | CDS | NM_020622 | ILMN_2789762 |
| Slc30a3 | Solute carrier family 30 (zinc transporter), member 3 | ENSMUSG00000029151 | 22784 | 8.12 | -0.147 | 0.000253807 | 4.73E-02 | -3.98 | chr5:31390678:31390689:-,chr5:31390427:31390464:- | CDS | NM_011773 | ILMN_2692938 |
| 9530068E07Rik | RIKEN cDNA 9530068E07 gene | ENSMUSG00000036275 | 67938 | 7.60 | -0.085 | 0.000254216 | 4.73E-02 | -3.98 | chr11:52216728:52216777:+ | CDS | NM_153117 | ILMN_2768859 |
| Eftud2 | Elongation factor Tu GTP binding domain containing 2 | ENSMUSG00000020929 | 12804 | 9.71 | -0.249 | 0.000258428 | 4.73E-02 | -3.98 | chr11:102700792:102700841:- | CDS | NM_011431 NM_001109995 | ILMN_2604097 |
| AI316807 | Expressed sequence AI316807 | ENSMUSG00000031534 | 52635 | 10.00 | -0.318 | 0.000260444 | 4.73E-02 | -3.97 | chr8:23573108:23573157:- | 3pUTR | NM_001012667 | ILMN_3160963 |
| Hbp1 | High mobility group box transcription factor 1 | ENSMUSG00000002996 | 230784 | 8.46 | -0.320 | 0.00026348 | 4.73E-02 | -3.97 | chr12:32615773:32615822:- | CDS | NM_177993 NM_153198 | ILMN_3092415 |
| Srp19 | Signal recognition particle 19 | ENSMUSG00000014504 | 230582 | 7.96 | 0.143 | 0.000265791 | 4.73E-02 | 3.97 | chr18:34491420:34491469:+ | CDS | NM_025527 | ILMN_2757224 |
| Prmt7 | Protein arginine N-methyltransferase 7 | ENSMUSG00000060098 | 214572 | 8.26 | -0.242 | 0.000268968 | 4.73E-02 | -3.96 | chr8:108761180:108761229:+ | CDS | NM_145404 | ILMN_1252819 |
| Pdcd5 | Programmed cell death 5 | ENSMUSG00000030417 | 100042424 | 7.82 | 0.141 | 0.000269866 | 4.73E-02 | 3.96 | chr7:36428662:36428711:- | CDS | XM_001478256 NM_019746 XM_001476181 | ILMN_2756658 |
| Smek2 | SMEK homolog 2, suppressor of mek1 (Dictyostelium) | ENSMUSG00000020463 | 68311 | 8.27 | 0.314 | 0.00027205 | 4.73E-02 | 3.96 | chr11:29119046:29119095:+ | 3pUTR | NM_134034 | ILMN_2907928 |
| Hcn2 | Hyperpolarization-activated, cyclic nucleotide-gated K+ 2 | ENSMUSG00000020331 | 13144 | 10.36 | 0.389 | 0.000274127 | 4.73E-02 | 3.96 | chr10:79198800:79198849:+ | 3pUTR | NM_008226 | ILMN_2672624 |
| Galnt2 | UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 2 | ENSMUSG00000031977 | 108902 | 7.79 | 0.124 | 0.000274756 | 4.73E-02 | 3.96 | chr8:126862454:126862469:+,chr8:126864719:126864752:+ | CDS | NM_139272 | ILMN_1242677 |
| Lpgat1 | Lysophosphatidylglycerol acyltransferase 1 | ENSMUSG00000026623 | 170755 | 7.63 | 0.082 | 0.000275105 | 4.73E-02 | 3.96 | chr1:193602405:193602454:+ | CDS | NM_172266 | ILMN_2720479 |
| Arpp19 | CAMP-regulated phosphoprotein 19 | ENSMUSG00000007656 | 59046 | 9.87 | 0.282 | 0.000275509 | 4.73E-02 | 3.95 | chr9:74907936:74907985:+ | 3pUTR | NM_021548 | ILMN_2674407 |
| Kcng4 | Potassium voltage-gated channel, subfamily G, member 4 | ENSMUSG00000045246 | 66921 | 8.02 | 0.152 | 0.000278033 | 4.73E-02 | 3.95 | chr8:122148119:122148168:- | 3pUTR | NM_025734 | ILMN_2895511 |
| 7.65 | -0.135 | 0.000278628 | 4.73E-02 | -3.95 | ILMN_2480596 | |||||||
| Eef1a1 | Eukaryotic translation elongation factor 1 alpha 1 | ENSMUSG00000037742 | 13627 | 14.87 | -0.286 | 0.000278681 | 4.73E-02 | -3.95 | chr9:78326765:78326775:-,chr9:78326627:78326665:- | CDS | NM_010106 | ILMN_2588052 |
| Carhsp1 | Calcium regulated heat stable protein 1 | 66411 | 9.32 | -0.152 | 0.000284903 | 4.73E-02 | -3.94 | chr16:8658946:8658995:- | 3pUTR | NM_025821 | ILMN_1225035 | |
| Nt5m | 5,3-nucleotidase, mitochondrial | ENSMUSG00000032615 | 215008 | 8.13 | 0.151 | 0.000285758 | 4.73E-02 | 3.94 | chr11:59689624:59689673:+ | 3pUTR | NM_134029 | ILMN_2987854 |
| Phkb | Phosphorylase kinase beta | ENSMUSG00000036879 | 106200 | 7.60 | -0.084 | 0.000290856 | 4.73E-02 | -3.94 | chr8:88584180:88584229:+ | 3pUTR | NM_199446 | ILMN_2807757 |
| Ncoa6 | Nuclear receptor coactivator 6 | ENSMUSG00000038369 | 21987 | 8.61 | -0.354 | 0.000295204 | 4.73E-02 | -3.93 | chr2:155231300:155231349:- | CDS | NM_019825 | ILMN_2749937 |
| Cox7b | Cytochrome c oxidase subunit VIIb | ENSMUSG00000031231 | 22311 | 7.86 | 0.222 | 0.000298824 | 4.73E-02 | 3.93 | chrX:103217157:103217206:+ | CDS | NM_025379 | ILMN_2671689 |
| Lpcat3 | Lysophosphatidylcholine acyltransferase 3 | ENSMUSG00000004270 | 66408 | 9.29 | -0.208 | 0.000300051 | 4.73E-02 | -3.93 | chr6:124653623:124653645:+,chr6:124653818:124653844:+ | CDS | NM_145130 | ILMN_2980858 |
| U83174 | Mus musculus ROSA 26 transcription 1 mRNA sequence. | 383348 | 7.83 | -0.178 | 0.000300954 | 4.73E-02 | -3.93 | chr6:113027069:113027118:- | 3pUTR | XM_489149* XM_924614* | ILMN_1215693 | |
| Lhx3 | LIM homeobox protein 3 | ENSMUSG00000026934 | 16871 | 7.66 | -0.096 | 0.000302811 | 4.73E-02 | -3.92 | chr2:26056071:26056120:- | 3pUTR | NM_001039653 | ILMN_2784382 |
| Nrp1 | Neuropilin 1 | ENSMUSG00000025810 | 74249 | 10.78 | 0.288 | 0.000305702 | 4.73E-02 | 3.92 | chr8:131027112:131027161:+ | 3pUTR | NM_008737 | ILMN_2669912 |
| Nipbl | Nipped-B homolog (Drosophila) | ENSMUSG00000022141 | 71175 | 9.69 | 0.640 | 0.000312201 | 4.73E-02 | 3.91 | chr15:8309179:8309228:- | CDS | NM_201232 NM_027707 | ILMN_1234693 |
| Usp28 | Ubiquitin specific peptidase 28 | ENSMUSG00000032267 | 69926 | 8.27 | -0.238 | 0.000312306 | 4.73E-02 | -3.91 | chr9:48850104:48850153:+ | 3pUTR | NM_175482 | ILMN_2828768 |
| Apoa1bp | Apolipoprotein A-I binding protein | ENSMUSG00000028070 | 14661 | 8.68 | 0.285 | 0.000312464 | 4.73E-02 | 3.91 | chr3:87860459:87860508:- | CDS | NM_144897 | ILMN_1240276 |
| Gm632 | MKIAA1196 protein (Fragment). | ENSMUSG00000000823 | 666200 | 10.75 | -0.337 | 0.00031502 | 4.73E-02 | -3.91 | chr2:181316912:181316961:- | 3pUTR | XM_990196 XM_977462 | ILMN_1219778 |
| Eng | Endoglin | ENSMUSG00000026814 | 16333 | 7.97 | 0.149 | 0.000317168 | 4.73E-02 | 3.91 | chr2:32537324:32537373:+ | 3pUTR | NM_007932 | ILMN_1229161 |
| Dalrd3 | DALR anticodon binding domain containing 3 | ENSMUSG00000019039 | 217151 | 9.66 | -0.286 | 0.000317563 | 4.73E-02 | -3.91 | chr9:108474567:108474616:+ | CDS | NM_026378 | ILMN_2644238 |
| 2900024O10Rik | RIKEN cDNA 2900024O10 gene | ENSMUSG00000051098 | 16578 | 9.51 | -0.266 | 0.000318218 | 4.73E-02 | -3.91 | chr13:81891692:81891741:+ | 3pUTR | NM_028372 | ILMN_3160416 |
| Gats | Opposite strand transcription unit to Stag3 | ENSMUSG00000015944 | 18201 | 9.54 | 0.293 | 0.000318732 | 4.73E-02 | 3.91 | chr5:134617378:134617427:+ | 3pUTR | NM_030719 | ILMN_3036708 |
| Emp2 | Epithelial membrane protein 2 | ENSMUSG00000022505 | 13731 | 8.39 | -0.212 | 0.000322528 | 4.73E-02 | -3.90 | chr16:10282072:10282121:- | 3pUTR | NM_007929 | ILMN_2598103 |
| Rpl15 | Ribosomal protein L15 | ENSMUSG00000080932 | 327958 | 13.31 | -0.235 | 0.000322815 | 4.73E-02 | -3.90 | chr14:19102467:19102516:- | CDS | XM_001472467 XM_001472443 XM_001472419 XM_001472396 XR_032689 XM_909622 XM_001480702 NM_025586 XM_001481032 XM_001481031 XM_001473843 XM_001473827 XM_001473807 XM_001473788 XM_001480293 | ILMN_1243415 |
| Sgtb | Small glutamine-rich tetratricopeptide repeat (TPR)-containing, beta | ENSMUSG00000042743 | 104086 | 8.71 | 0.355 | 0.000326026 | 4.73E-02 | 3.90 | chr13:104930948:104930997:+ | 3pUTR | NM_144838 | ILMN_1219768 |
| 2810441K11Rik | hypothetical protein LOC68642 | ENSMUSG00000024667 | 7.61 | 0.061 | 0.000326851 | 4.73E-02 | 3.90 | chr19:10625448:10625486:-,chr19:10614298:10614308:- | 3pUTR | NM_026798 | ILMN_2545299 | |
| Commd7 | COMM domain containing 7 | ENSMUSG00000056941 | 109349 | 8.86 | 0.202 | 0.000331148 | 4.73E-02 | 3.89 | chr2:153444612:153444661:- | 3pUTR | NM_133850 XM_905642 XM_979437 | ILMN_1229682 |
| Ddx5 | DEAD (Asp-Glu-Ala-Asp) box polypeptide 5 | ENSMUSG00000082454 | 13207 | 12.56 | -0.298 | 0.000333978 | 4.73E-02 | -3.89 | chr11:106643188:106643234:- | CDS | XR_031177 XR_031980 | ILMN_1253539 |
| Asphd2 | Aspartate beta-hydroxylase domain containing 2 | ENSMUSG00000029348 | 72898 | 11.69 | -0.263 | 0.000336186 | 4.73E-02 | -3.89 | chr5:112814553:112814602:- | 3pUTR | NM_028386 | ILMN_3162247 |
| Mdga2 | MAM domain containing glycosylphosphatidylinositol anchor 2 | ENSMUSG00000034912 | 8.34 | 0.250 | 0.000338288 | 4.73E-02 | 3.89 | chr12:67574189:67574215:-,chr12:67572568:67572590:- | CDS | NM_207010 | ILMN_2847711 | |
| Ncam1 | Neural cell adhesion molecule 1 | 73072 | 8.04 | -0.223 | 0.000338844 | 4.73E-02 | -3.89 | chr9:49450189:49450238:- | 3pUTR | ILMN_1254093 | ||
| Kcng4 | Potassium voltage-gated channel, subfamily G, member 4 | ENSMUSG00000045246 | 235636 | 7.86 | 0.114 | 0.000345633 | 4.73E-02 | 3.88 | chr8:122147783:122147832:- | 3pUTR | NM_025734 | ILMN_1233788 |
| mt-Nd4 | NADH dehydrogenase subunit 4L (EC 1.6.5.3) (Activated spleen cDNA, RIKEN full-length enriched library, clone:F830212L04 product:NADH dehydrogenase 4, mitochondrial, full insert sequence) (3 days neonate thymus cDNA, RIKEN full-length enriched library, clone:A630072D15 product:NADH dehydrogenase 4, mitochondrial, full insert sequence) (Lung RCB-0558 LLC cDNA, RIKEN full-length enriched library, clone:G730042M17 product:NADH dehydrogenase 4, mitochondrial, full insert sequence) (Adult male corpora quadrigemina cDNA, RIKEN full- length enriched library, clone:B230333I23 product:NADH dehydrogenase 4, mitochondrial, full insert sequence). | ENSMUSG00000067736 | 12558 | 9.22 | 0.976 | 0.000347113 | 4.73E-02 | 3.88 | chrM:10117:10166:+ | CDS | ILMN_2419660 | |
| Thsd7b | Thrombospondin, type I, domain containing 7B | ENSMUSG00000042581 | 14450 | 8.70 | -0.246 | 0.000348235 | 4.73E-02 | -3.88 | chr1:132115331:132115380:+ | 3pUTR | NM_172485 | ILMN_2949380 |
| Asns | Asparagine synthetase | ENSMUSG00000029752 | 231290 | 9.92 | 0.183 | 0.000349115 | 4.73E-02 | 3.88 | chr6:7625993:7625996:-,chr6:7625479:7625524:- | CDS | NM_012055 | ILMN_3006123 |
| Prl2c3 | Prolactin family 2, subfamily c, member 3 | ENSMUSG00000056457 | 497106 | 7.59 | 0.079 | 0.000349843 | 4.73E-02 | 3.88 | chr13:12892287:12892336:- | 5pUTR | NM_011118 | ILMN_2875585 |
| Fam65b | Family with sequence similarity 65, member B | ENSMUSG00000036006 | 193385 | 9.68 | -0.176 | 0.000350327 | 4.73E-02 | -3.88 | chr13:24825086:24825135:+ | 3pUTR | NM_029679 | ILMN_1255560 |
| Nrxn3 | Neurexin III | ENSMUSG00000066392 | 98733 | 9.56 | 0.664 | 0.000353555 | 4.73E-02 | 3.87 | chr12:91571391:91571440:+ | 3pUTR | NM_172544 | ILMN_2763404 |
| Fem1b | Feminization 1 homolog b (C. elegans) | ENSMUSG00000032244 | 72999 | 11.16 | -0.168 | 0.000357194 | 4.73E-02 | -3.87 | chr9:62639952:62640001:- | 3pUTR | NM_010193 | ILMN_1230162 |
| Cdc7 | Cell division cycle 7 (S. cerevisiae) | ENSMUSG00000029283 | 258427 | 9.34 | -0.239 | 0.000358118 | 4.73E-02 | -3.87 | chr5:107413236:107413285:+ | 3pUTR | NM_009863 | ILMN_1238374 |
| Tex11 | Testis expressed gene 11 | ENSMUSG00000009670 | 237761 | 7.62 | 0.081 | 0.000358881 | 4.73E-02 | 3.87 | chrX:98034079:98034128:- | 3pUTR | NM_031384 | ILMN_2599470 |
| Zfp292 | Zinc finger protein 292 | ENSMUSG00000039967 | 66905 | 8.33 | 0.375 | 0.000359144 | 4.73E-02 | 3.87 | chr4:34752198:34752247:- | CDS | NM_013889 | ILMN_1243910 |
| Ctdp1 | CTD (carboxy-terminal domain, RNA polymerase II, polypeptide A) phosphatase, subunit 1 | ENSMUSG00000033323 | 93679 | 7.54 | 0.061 | 0.000359225 | 4.73E-02 | 3.87 | chr18:80605201:80605250:- | 3pUTR | NM_026295 | ILMN_2650492 |
| Cpe | Carboxypeptidase E | ENSMUSG00000037852 | 13135 | 14.64 | -0.339 | 0.000365669 | 4.73E-02 | -3.86 | chr8:67073713:67073762:- | CDS | NM_013494 XM_001002936 | ILMN_1228232 |
| Thoc7 | THO complex 7 homolog (Drosophila) | ENSMUSG00000053453 | 20822 | 7.97 | 0.211 | 0.000365696 | 4.73E-02 | 3.86 | chr14:14787053:14787100:-,chr14:14786051:14786052:- | CDS | NM_001013578 NM_025435 XM_001473442 XM_001473268 | ILMN_2600729 |
| Chd9 | Chromodomain helicase DNA binding protein 9 | ENSMUSG00000056608 | 381628 | 7.61 | -0.092 | 0.00036841 | 4.73E-02 | -3.86 | chr8:93510011:93510060:+ | CDS | NM_177224 | ILMN_2600528 |
| S100a10 | S100 calcium binding protein A10 (calpactin) | ENSMUSG00000041959 | 20194 | 7.86 | 0.204 | 0.000371081 | 4.73E-02 | 3.86 | chr3:93368411:93368460:+ | 3pUTR | NM_009112 | ILMN_1256702 |
| 9030025P20Rik | RIKEN cDNA 9030025P20 gene | ENSMUSG00000036552 | 18096 | 8.42 | -0.185 | 0.000375045 | 4.73E-02 | -3.85 | chr17:15198084:15198133:+ | CDS | NM_001034891 NM_001039552 | ILMN_2716783 |
| Akap1 | A kinase (PRKA) anchor protein 1 | ENSMUSG00000018428 | 13508 | 7.58 | 0.066 | 0.000377879 | 4.73E-02 | 3.85 | chr11:88725839:88725888:- | 5pUTR | NM_001042541 | ILMN_3068062 |
| abParts | Parts of antibodies, mostly variable regions. | ENSMUSG00000076591 | 216345 | 7.60 | -0.074 | 0.000380078 | 4.73E-02 | -3.85 | chr6:70337184:70337200:-,chr6:70336949:70336981:- | 3pUTR | ILMN_2462709 | |
| Rbm12 | RNA binding motif protein 12 | ENSMUSG00000038171 | 56219 | 8.18 | -0.113 | 0.000388928 | 4.73E-02 | -3.84 | chr2:155921168:155921217:- | CDS | NM_170598 NM_029397 | ILMN_3104139 |
| Cacnb3 | Calcium channel, voltage-dependent, beta 3 subunit | ENSMUSG00000003352 | 328329 | 8.50 | -0.233 | 0.000396235 | 4.73E-02 | -3.84 | chr15:98473935:98473984:+ | CDS | NM_001044741 NM_007581 | ILMN_2702286 |
| Ltk | Leukocyte tyrosine kinase | ENSMUSG00000027297 | 69922 | 8.37 | -0.187 | 0.000396467 | 4.73E-02 | -3.84 | chr2:119577207:119577256:- | 3pUTR | NM_206942 NM_008523 NM_206941 NM_203345 | ILMN_1219860 |
| Pcdhb14 | Protocadherin beta 14 | ENSMUSG00000044043 | 71026 | 7.80 | 0.115 | 0.00039748 | 4.73E-02 | 3.83 | chr18:37609963:37610012:+ | 3pUTR | NM_053139 | ILMN_2607000 |
| Rab23 | RAB23, member RAS oncogene family | ENSMUSG00000004768 | 17207 | 8.04 | -0.177 | 0.000397783 | 4.73E-02 | -3.83 | chr1:33798015:33798064:+ | 3pUTR | NM_008999 | ILMN_2687881 |
| Slc5a5 | Solute carrier family 5 (sodium iodide symporter), member 5 | ENSMUSG00000000792 | 23821 | 9.24 | -0.141 | 0.000398304 | 4.73E-02 | -3.83 | chr8:73406944:73406993:- | 3pUTR | NM_053248 | ILMN_1245214 |
| Ctnnd2 | Catenin (cadherin associated protein), delta 2 | ENSMUSG00000022240 | 232975 | 8.92 | -0.440 | 0.000399149 | 4.73E-02 | -3.83 | chr15:30896661:30896710:+ | CDS | XR_032172 NM_008729 | ILMN_2652527 |
| Rgs3 | Regulator of G-protein signaling 3 | ENSMUSG00000059810 | 242646 | 7.64 | -0.092 | 0.00039942 | 4.73E-02 | -3.83 | chr4:62267576:62267625:+ | CDS | NM_019492 | ILMN_2610679 |
| Ano9 | Anoctamin 9 | ENSMUSG00000054662 | 68035 | 7.59 | 0.077 | 0.000402659 | 4.73E-02 | 3.83 | chr7:148287144:148287193:- | 3pUTR | NM_178381 | ILMN_1242413 |
| C330023M02Rik | RIKEN cDNA C330023M02 gene | ENSMUSG00000042719 | 22770 | 9.39 | 0.407 | 0.000403643 | 4.73E-02 | 3.83 | chr5:121887636:121887685:+ | CDS | NM_172722 | ILMN_1228917 |
| Olfr203 | Olfactory receptor 203 | ENSMUSG00000068182 | 20019 | 7.57 | 0.078 | 0.00040471 | 4.73E-02 | 3.83 | chr16:59302997:59303046:+ | CDS | NM_146486 | ILMN_1214508 |
| Vamp2 | Vesicle-associated membrane protein 2 | ENSMUSG00000020894 | 68197 | 13.01 | 0.192 | 0.000411184 | 4.73E-02 | 3.82 | chr11:68905817:68905866:+ | 3pUTR | NM_009497 | ILMN_2424912 |
| Hist1h2an | Histone cluster 1, H2an | ENSMUSG00000069309 | 14263 | 8.39 | 0.441 | 0.000415867 | 4.73E-02 | 3.82 | chr13:21878698:21878747:- | 3pUTR | NM_178184 | ILMN_1248830 |
| Ddx6 | DEAD (Asp-Glu-Ala-Asp) box polypeptide 6 | ENSMUSG00000032097 | 66419 | 7.85 | 0.220 | 0.000422588 | 4.73E-02 | 3.81 | chr9:44443908:44443948:+,chr9:44444572:44444580:+ | CDS | NM_181324 NM_001110826 NM_007841 | ILMN_3114632 |
| Gpatch2 | G patch domain containing 2 | ENSMUSG00000039210 | 67769 | 9.10 | -0.122 | 0.000422854 | 4.73E-02 | -3.81 | chr1:189175118:189175167:+ | 3pUTR | NM_026367 | ILMN_2841032 |
| Ptp4a2 | Protein tyrosine phosphatase 4a2 | ENSMUSG00000028788 | 67695 | 8.79 | -0.354 | 0.000427844 | 4.73E-02 | -3.81 | chr4:129525764:129525813:+ | 3pUTR | NM_008974 | ILMN_1243291 |
| Slitrk4 | SLIT and NTRK-like family, member 4 | ENSMUSG00000046699 | 20742 | 9.26 | 0.576 | 0.000429398 | 4.73E-02 | 3.81 | chrX:61522862:61522911:- | 3pUTR | NM_178740 | ILMN_2715848 |
| Akap7 | A kinase (PRKA) anchor protein 7 | ENSMUSG00000039166 | 258486 | 9.76 | -0.194 | 0.000432535 | 4.73E-02 | -3.81 | chr10:24889980:24890029:- | 3pUTR | NM_018747 | ILMN_2944413 |
| Arfgap2 | ADP-ribosylation factor GTPase activating protein 2 | ENSMUSG00000027255 | 9.12 | -0.231 | 0.000433582 | 4.73E-02 | -3.81 | chr2:91116851:91116900:+ | 3pUTR | NM_023854 | ILMN_1241481 | |
| Vegfc | Vascular endothelial growth factor C | ENSMUSG00000031520 | 15560 | 7.96 | 0.163 | 0.000438865 | 4.73E-02 | 3.80 | chr8:55271361:55271410:+ | CDS | NM_009506 | ILMN_2486573 |
| Elfn2 | Leucine rich repeat and fibronectin type III, extracellular 2 | ENSMUSG00000043460 | 50876 | 9.44 | 0.476 | 0.000439324 | 4.73E-02 | 3.80 | chr15:78500612:78500661:- | 3pUTR | NM_183141 | ILMN_1239570 |
Supplementary Table 2. Antibodies used in WB, IP, IHC and IF assays
List of antibodies used in biochemical assays. Antibody: antibody target; Assay: assay in which the antibody was used; Dilution/concentration: Dilution and/or concentration used for each antibody; Company and Catalogue Number: company name and catalogue number for commercially available antibodies.
| Antibody | Assay | Dilution/concentration | Company | Catalogue Number |
|---|---|---|---|---|
| b-catenin | WB | 1/2000 | Cell signaling | 9582 |
| CRMP2 | WB-IP | 1/1000 - 1ug/100ul | Cell Signaling | 9393 |
| CRMP2 pT514 | WB | 1/1000 | Cell Signaling | 9397 |
| DISC1 | WB-IP | 1ug/ml- 1ug/100ul | Invitrogen | 40-6800 |
| DISC1 | WB-IP | 1ug/ml- 1ug/100ul | Invitrogen | 40-6900 |
| DISC1 | WB | 1/500 | Gift from Nick Brandon | n/a |
| Doublecortin | IHC | 1/250 | Cell signaling | 4604 |
| GluR1 | WB-IP | 1/5000 - 1ug/100ul | Millipore | AB1504 |
| GluR1 | WB-IP | 1/2000 - 1ug/100ul | Millipore | 05-855 |
| GlurR1 | WB | 1/1000 | Santa Cruz | sc-7609 |
| GSK3-b | WB-IP | 1/1000 | Cell Signaling | 9332 |
| Ki67 | IF | 1/300 | Abcam | ab15580 |
| NeuroD1 | WB | 1/1000 | Cell signaling | 4373 |
| NeuroD1 | IF | 1/500 | Abcam | ab60704 |
| NeuroD1 pS274 | WB | 1/1000 | Abcam | ab78900 |
| NR1 | WB-IP | 1/1000 - 1ug/100ul | Invitrogen | 32-0500 |
| NR1 | IP | 1ug/100ul | Created in house | n/a |
| NR1 | WB | 1ug/ml | Millipore | 06-311 |
| NR2B | WB | 1ug/ml | Millipore | 05-920 |
| NR2B | WB | 1/2000 - 1ug/100ul | Millipore | 06-600 |
| PSD95 | WB | 1/10000 | Thermo | MA1-045 |
| PSD95 pT19 | WB | 1ug/ml | Abcam | ab16496 |
| PSD95 pS418 | WB | 1ug/ml | Abcam | ab16493 |
| Tau pS396 | WB | 1/1000 | Gift from Michel Goedert | n/a |
| TNiK | WB-IP | 0.5ug/ml- 0.25ug/100ul | BD biosciences | 612250 |
| TNiK | WB | 1/10000 | Santa Cruz | sc-100206 |
| TNiK | WB | 1ug/ml- 0.5ug/100ul | Sigma | HPA012128 |
| TNiK | IP | 1ug/100ul | Thermo | PA1-20639 |
| TNiK pS735 | WB-IP | 1ug/ml- 1ug/100ul | Santa Cruz | sc-130221 |
| Tubulin | WB | 1ug/ml | Millipore | 05-559 |