Automated design of genomic Southern blot probes
Mike DR Croning, David G Fricker, Noboru H Komiyama and Seth GN Grant
Design Case - ProbeDesign47 - back to list
Summary
./analyse_probe_search_results : Wed Sep 21 13:54:29 2005
Connected to host: host, as user: user Database : southern_blot_designs Fetched probe design: ProbeDesign47 assembly : NCBIM34 bias : 3prime chromosome : 8 id : 29 strand : 1 ------------- Design window length: 4712bp All jobs for probe design are successful - total: 81 Fetched conf : Exonerate mouse_NCBIM34 (id: 1) Genome file(s) at: /blastdb/Mouse/NCBIM34/softmasked_dusted Fetched 1611 putative probes to analyse Selection criteria to rank probes: Minimum score ratio : 10 (i.e. self-hit must score at least 10 times higher than the next best hit) Maximum percent repetitive bases: 5 Favour probes to the 3prime end of design window ------------------------ Putative probes without hits : 0 Putative probes with single_self_hit : 1611/1611 Putative probes exceeding selection criteria: 210/1611 PASSED (GREEN) AND FAILED PUTATIVE PROBES IN THE DESIGN WINDOW --------------------------------------------------------------
Image
Unique (yellow), passed (green) and failed putative probes in the design window
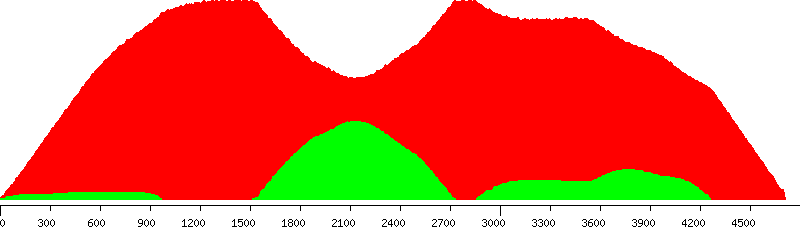
Unique putative probes
No probes
Putative probes
| Putative probe ID | Length | Score ratio | Actual | %rep DNA | Distance | Overall |
|---|---|---|---|---|---|---|
| 30780 | 1150 | 33.2 | 5750 | 3.1 | 1995 | 27.0 |
| 30845 | 1100 | 31.8 | 5500 | 2.5 | 2035 | 26.7 |
| 30913 | 1050 | 30.3 | 5250 | 2.7 | 2028 | 25.0 |
| 30914 | 1050 | 30.3 | 5250 | 2.7 | 2080 | 25.0 |
| 30986 | 1000 | 28.9 | 5000 | 2.8 | 2050 | 23.3 |
| 30987 | 1000 | 28.9 | 5000 | 2.8 | 2100 | 23.3 |
| 30988 | 1000 | 28.9 | 5000 | 2.8 | 2150 | 23.3 |
| 30985 | 1000 | 28.9 | 5000 | 3.1 | 2000 | 22.7 |
| 30844 | 1100 | 31.8 | 5500 | 4.6 | 1980 | 22.5 |
| 31151 | 900 | 26.0 | 4500 | 2.2 | 2250 | 21.6 |
| 31063 | 950 | 27.5 | 4750 | 2.9 | 2021 | 21.6 |
| 31064 | 950 | 27.5 | 4750 | 2.9 | 2068 | 21.6 |
| 31065 | 950 | 27.5 | 4750 | 2.9 | 2115 | 21.6 |
| 31066 | 950 | 27.5 | 4750 | 2.9 | 2162 | 21.6 |
| 31067 | 950 | 27.5 | 4750 | 2.9 | 2209 | 21.6 |
| 30915 | 1050 | 30.3 | 5250 | 4.5 | 2132 | 21.4 |
| 31328 | 800 | 23.8 | 4000 | 1.4 | 2000 | 21.1 |
| 30846 | 1100 | 30.9 | 5500 | 5.0 | 2090 | 20.9 |
| 31431 | 750 | 22.3 | 3750 | 1.1 | 2035 | 20.2 |
| 31240 | 850 | 24.6 | 4250 | 2.4 | 2268 | 19.9 |
| 31241 | 850 | 24.6 | 4250 | 2.4 | 2310 | 19.9 |
| 31146 | 900 | 26.0 | 4500 | 3.1 | 2025 | 19.8 |
| 31147 | 900 | 26.0 | 4500 | 3.1 | 2070 | 19.8 |
| 31148 | 900 | 26.0 | 4500 | 3.1 | 2115 | 19.8 |
| 31149 | 900 | 26.0 | 4500 | 3.1 | 2160 | 19.8 |
| 31150 | 900 | 26.0 | 4500 | 3.1 | 2205 | 19.8 |
| 31430 | 750 | 22.3 | 3750 | 1.7 | 1998 | 18.9 |
| 31100 | 950 | 26.7 | 4750 | 4.0 | 3760 | 18.7 |
| 31542 | 700 | 20.8 | 3500 | 1.1 | 2030 | 18.5 |
| 31543 | 700 | 20.8 | 3500 | 1.1 | 2065 | 18.5 |
| 31544 | 700 | 20.8 | 3500 | 1.1 | 2100 | 18.5 |
| 31432 | 750 | 22.3 | 3750 | 2.0 | 2072 | 18.3 |
| 31335 | 800 | 23.1 | 4000 | 2.5 | 2280 | 18.1 |
| 31336 | 800 | 23.1 | 4000 | 2.5 | 2320 | 18.1 |
| 31337 | 800 | 23.1 | 4000 | 2.5 | 2360 | 18.1 |
| 31662 | 650 | 19.3 | 3250 | 1.2 | 2016 | 16.9 |
| 31663 | 650 | 19.3 | 3250 | 1.2 | 2048 | 16.9 |
| 31664 | 650 | 19.3 | 3250 | 1.2 | 2080 | 16.9 |
| 31665 | 650 | 19.3 | 3250 | 1.2 | 2112 | 16.9 |
| 31666 | 650 | 19.3 | 3250 | 1.2 | 2144 | 16.9 |
| 31437 | 750 | 21.7 | 3750 | 2.7 | 2257 | 16.3 |
| 31438 | 750 | 21.7 | 3750 | 2.7 | 2294 | 16.3 |
| 31439 | 750 | 21.7 | 3750 | 2.7 | 2331 | 16.3 |
| 31440 | 750 | 21.7 | 3750 | 2.7 | 2368 | 16.3 |
| 31441 | 750 | 21.7 | 3750 | 2.7 | 2405 | 16.3 |
| 31541 | 700 | 20.8 | 3500 | 2.3 | 1995 | 16.3 |
| 31851 | 600 | 17.4 | 3000 | 1.0 | 3750 | 15.4 |
| 32003 | 550 | 16.4 | 2750 | 0.5 | 3753 | 15.3 |
| 31793 | 600 | 17.9 | 3000 | 1.3 | 2010 | 15.2 |
| 31794 | 600 | 17.9 | 3000 | 1.3 | 2040 | 15.2 |
| 31795 | 600 | 17.9 | 3000 | 1.3 | 2070 | 15.2 |
| 31796 | 600 | 17.9 | 3000 | 1.3 | 2100 | 15.2 |
| 31797 | 600 | 17.9 | 3000 | 1.3 | 2130 | 15.2 |
| 31798 | 600 | 17.9 | 3000 | 1.3 | 2160 | 15.2 |
| 31799 | 600 | 17.9 | 3000 | 1.3 | 2190 | 15.2 |
| 31549 | 700 | 20.2 | 3500 | 2.9 | 2275 | 14.5 |
| 31550 | 700 | 20.2 | 3500 | 2.9 | 2310 | 14.5 |
| 31551 | 700 | 20.2 | 3500 | 2.9 | 2345 | 14.5 |
| 31552 | 700 | 20.2 | 3500 | 2.9 | 2380 | 14.5 |
| 31553 | 700 | 20.2 | 3500 | 2.9 | 2415 | 14.5 |
| 31554 | 700 | 20.2 | 3500 | 2.9 | 2450 | 14.5 |
| 32072 | 500 | 15.8 | 2500 | 0.8 | 1325 | 14.2 |
| 31667 | 650 | 19.3 | 3250 | 2.9 | 2176 | 13.5 |
| 31939 | 550 | 16.4 | 2750 | 1.5 | 2025 | 13.5 |
| 31940 | 550 | 16.4 | 2750 | 1.5 | 2052 | 13.5 |
| 31941 | 550 | 16.4 | 2750 | 1.5 | 2079 | 13.5 |
| 31942 | 550 | 16.4 | 2750 | 1.5 | 2106 | 13.5 |
| 31943 | 550 | 16.4 | 2750 | 1.5 | 2133 | 13.5 |
| 31944 | 550 | 16.4 | 2750 | 1.5 | 2160 | 13.5 |
| 31945 | 550 | 16.4 | 2750 | 1.5 | 2187 | 13.5 |
| 31670 | 650 | 18.8 | 3250 | 3.1 | 2272 | 12.6 |
| 31671 | 650 | 18.8 | 3250 | 3.1 | 2304 | 12.6 |
| 31672 | 650 | 18.8 | 3250 | 3.1 | 2336 | 12.6 |
| 31673 | 650 | 18.8 | 3250 | 3.1 | 2368 | 12.6 |
| 31674 | 650 | 18.8 | 3250 | 3.1 | 2400 | 12.6 |
| 31675 | 650 | 18.8 | 3250 | 3.1 | 2432 | 12.6 |
| 31676 | 650 | 18.8 | 3250 | 3.1 | 2464 | 12.6 |
| 31677 | 650 | 18.8 | 3250 | 3.1 | 2496 | 12.6 |
| 32107 | 500 | 15.4 | 2500 | 1.6 | 2200 | 12.2 |
| 32169 | 500 | 14.5 | 2500 | 1.2 | 3750 | 12.1 |
| 31716 | 650 | 16.8 | 3250 | 2.5 | 3744 | 11.9 |
| 32073 | 500 | 15.8 | 2500 | 2.0 | 1350 | 11.8 |
| 32100 | 500 | 14.9 | 2500 | 1.6 | 2025 | 11.7 |
| 32101 | 500 | 14.9 | 2500 | 1.6 | 2050 | 11.7 |
| 32102 | 500 | 14.9 | 2500 | 1.6 | 2075 | 11.7 |
| 32103 | 500 | 14.9 | 2500 | 1.6 | 2100 | 11.7 |
| 32104 | 500 | 14.9 | 2500 | 1.6 | 2125 | 11.7 |
| 32105 | 500 | 14.9 | 2500 | 1.6 | 2150 | 11.7 |
| 32106 | 500 | 14.9 | 2500 | 1.6 | 2175 | 11.7 |
| 31938 | 550 | 16.4 | 2750 | 2.4 | 1998 | 11.6 |
| 31661 | 650 | 19.3 | 3250 | 4.2 | 1984 | 11.0 |
| 31811 | 600 | 17.3 | 3000 | 3.3 | 2550 | 10.7 |
| 31613 | 650 | 17.0 | 3250 | 3.2 | 448 | 10.6 |
| 31614 | 650 | 17.0 | 3250 | 3.2 | 480 | 10.6 |
| 32099 | 500 | 14.9 | 2500 | 2.2 | 2000 | 10.5 |
| 31742 | 600 | 15.7 | 3000 | 3.5 | 480 | 8.7 |
| 31743 | 600 | 15.7 | 3000 | 3.5 | 510 | 8.7 |
| 31744 | 600 | 14.0 | 3000 | 3.5 | 540 | 7.0 |
| 31881 | 550 | 14.4 | 2750 | 3.8 | 459 | 6.8 |
| 31882 | 550 | 14.4 | 2750 | 3.8 | 486 | 6.8 |
| 31883 | 550 | 14.4 | 2750 | 3.8 | 513 | 6.8 |
| 31884 | 550 | 14.4 | 2750 | 3.8 | 540 | 6.8 |
| 31885 | 550 | 14.4 | 2750 | 3.8 | 567 | 6.8 |
| 32125 | 500 | 14.5 | 2500 | 4.0 | 2650 | 6.5 |
| 31961 | 550 | 15.9 | 2750 | 4.7 | 2619 | 6.4 |
| 30584 | 1300 | 14.8 | 6500 | 4.2 | 455 | 6.3 |
| 30585 | 1300 | 14.8 | 6500 | 4.2 | 520 | 6.3 |
| 31497 | 700 | 12.1 | 3500 | 3.0 | 455 | 6.1 |
| 30889 | 1050 | 11.9 | 5250 | 3.2 | 780 | 5.5 |
| 32038 | 500 | 13.1 | 2500 | 4.2 | 475 | 4.7 |
| 32039 | 500 | 13.1 | 2500 | 4.2 | 500 | 4.7 |
| 32040 | 500 | 13.1 | 2500 | 4.2 | 525 | 4.7 |
| 32041 | 500 | 13.1 | 2500 | 4.2 | 550 | 4.7 |
| 32042 | 500 | 13.1 | 2500 | 4.2 | 575 | 4.7 |
| 32043 | 500 | 13.1 | 2500 | 4.2 | 600 | 4.7 |
| 32044 | 500 | 13.1 | 2500 | 4.2 | 625 | 4.7 |
| 30961 | 1000 | 11.4 | 5000 | 3.4 | 800 | 4.6 |
| 30960 | 1000 | 11.4 | 5000 | 3.6 | 750 | 4.2 |
| 31886 | 550 | 11.8 | 2750 | 3.8 | 594 | 4.1 |
| 31036 | 950 | 10.8 | 4750 | 3.6 | 752 | 3.6 |
| 31037 | 950 | 10.8 | 4750 | 3.6 | 799 | 3.6 |
| 31038 | 950 | 10.8 | 4750 | 3.6 | 846 | 3.6 |
| 31615 | 650 | 10.0 | 3250 | 3.2 | 512 | 3.6 |
| 30757 | 1150 | 13.1 | 5750 | 4.8 | 684 | 3.5 |
| 31039 | 950 | 10.8 | 4750 | 3.9 | 893 | 3.0 |
| 31118 | 900 | 10.2 | 4500 | 3.8 | 765 | 2.7 |
| 31119 | 900 | 10.2 | 4500 | 3.8 | 810 | 2.7 |
| 31120 | 900 | 10.2 | 4500 | 3.8 | 855 | 2.7 |
| 31121 | 900 | 10.2 | 4500 | 3.8 | 900 | 2.7 |
| 30962 | 1000 | 11.4 | 5000 | 4.4 | 850 | 2.6 |
| 31122 | 900 | 10.2 | 4500 | 4.3 | 945 | 1.6 |
Overall score = score_ratio - 2 * repetitive DNA content
Redundant probes
| Putative probe ID | Length | Score ratio | Actual | %rep DNA | Distance | Overall |
|---|---|---|---|---|---|---|
| 31234 | 850 | 24.9 | 4250 | 3.3 | 2016 | 18.3 |
| 31235 | 850 | 24.6 | 4250 | 3.3 | 2058 | 18.0 |
| 31236 | 850 | 24.6 | 4250 | 3.3 | 2100 | 18.0 |
| 31237 | 850 | 24.6 | 4250 | 3.3 | 2142 | 18.0 |
| 31238 | 850 | 24.6 | 4250 | 3.3 | 2184 | 18.0 |
| 31239 | 850 | 24.6 | 4250 | 3.3 | 2226 | 18.0 |
| 31185 | 900 | 25.3 | 4500 | 4.2 | 3780 | 16.8 |
| 31329 | 800 | 23.8 | 4000 | 3.5 | 2040 | 16.8 |
| 31330 | 800 | 23.1 | 4000 | 3.5 | 2080 | 16.1 |
| 31331 | 800 | 23.1 | 4000 | 3.5 | 2120 | 16.1 |
| 31332 | 800 | 23.1 | 4000 | 3.5 | 2160 | 16.1 |
| 31333 | 800 | 23.1 | 4000 | 3.5 | 2200 | 16.1 |
| 31334 | 800 | 23.1 | 4000 | 3.5 | 2240 | 16.1 |
| 31276 | 850 | 23.9 | 4250 | 4.5 | 3780 | 14.9 |
| 31277 | 850 | 23.9 | 4250 | 4.5 | 3822 | 14.9 |
| 31433 | 750 | 21.9 | 3750 | 3.7 | 2109 | 14.5 |
| 31434 | 750 | 21.7 | 3750 | 3.7 | 2146 | 14.2 |
| 31435 | 750 | 21.7 | 3750 | 3.7 | 2183 | 14.2 |
| 31436 | 750 | 21.7 | 3750 | 3.7 | 2220 | 14.2 |
| 31372 | 800 | 22.5 | 4000 | 4.8 | 3760 | 13.0 |
| 31373 | 800 | 22.5 | 4000 | 4.8 | 3800 | 13.0 |
| 31374 | 800 | 22.5 | 4000 | 4.8 | 3840 | 13.0 |
| 31375 | 800 | 22.5 | 4000 | 4.8 | 3880 | 13.0 |
| 31545 | 700 | 20.8 | 3500 | 4.0 | 2135 | 12.8 |
| 31546 | 700 | 20.2 | 3500 | 4.0 | 2170 | 12.2 |
| 31547 | 700 | 20.2 | 3500 | 4.0 | 2205 | 12.2 |
| 31548 | 700 | 20.2 | 3500 | 4.0 | 2240 | 12.2 |
| 31801 | 600 | 17.9 | 3000 | 3.3 | 2250 | 11.2 |
| 31802 | 600 | 17.3 | 3000 | 3.3 | 2280 | 10.7 |
| 31803 | 600 | 17.3 | 3000 | 3.3 | 2310 | 10.7 |
| 31804 | 600 | 17.3 | 3000 | 3.3 | 2340 | 10.7 |
| 31805 | 600 | 17.3 | 3000 | 3.3 | 2370 | 10.7 |
| 31806 | 600 | 17.3 | 3000 | 3.3 | 2400 | 10.7 |
| 31807 | 600 | 17.3 | 3000 | 3.3 | 2430 | 10.7 |
| 31808 | 600 | 17.3 | 3000 | 3.3 | 2460 | 10.7 |
| 31809 | 600 | 17.3 | 3000 | 3.3 | 2490 | 10.7 |
| 31810 | 600 | 17.3 | 3000 | 3.3 | 2520 | 10.7 |
| 31668 | 650 | 19.0 | 3250 | 4.3 | 2208 | 10.4 |
| 31949 | 550 | 17.6 | 2750 | 3.6 | 2295 | 10.4 |
| 31669 | 650 | 18.8 | 3250 | 4.3 | 2240 | 10.2 |
| 31741 | 600 | 15.7 | 3000 | 3.5 | 450 | 8.7 |
| 31950 | 550 | 15.9 | 2750 | 3.6 | 2322 | 8.6 |
| 31951 | 550 | 15.9 | 2750 | 3.6 | 2349 | 8.6 |
| 31952 | 550 | 15.9 | 2750 | 3.6 | 2376 | 8.6 |
| 31953 | 550 | 15.9 | 2750 | 3.6 | 2403 | 8.6 |
| 31954 | 550 | 15.9 | 2750 | 3.6 | 2430 | 8.6 |
| 31955 | 550 | 15.9 | 2750 | 3.6 | 2457 | 8.6 |
| 31956 | 550 | 15.9 | 2750 | 3.6 | 2484 | 8.6 |
| 31957 | 550 | 15.9 | 2750 | 3.6 | 2511 | 8.6 |
| 31958 | 550 | 15.9 | 2750 | 3.6 | 2538 | 8.6 |
| 31959 | 550 | 15.9 | 2750 | 3.6 | 2565 | 8.6 |
| 31960 | 550 | 15.9 | 2750 | 3.6 | 2592 | 8.6 |
| 32113 | 500 | 14.9 | 2500 | 4.0 | 2350 | 6.9 |
| 32114 | 500 | 14.5 | 2500 | 4.0 | 2375 | 6.5 |
| 32115 | 500 | 14.5 | 2500 | 4.0 | 2400 | 6.5 |
| 32116 | 500 | 14.5 | 2500 | 4.0 | 2425 | 6.5 |
| 32117 | 500 | 14.5 | 2500 | 4.0 | 2450 | 6.5 |
| 32118 | 500 | 14.5 | 2500 | 4.0 | 2475 | 6.5 |
| 32119 | 500 | 14.5 | 2500 | 4.0 | 2500 | 6.5 |
| 32120 | 500 | 14.5 | 2500 | 4.0 | 2525 | 6.5 |
| 32121 | 500 | 14.5 | 2500 | 4.0 | 2550 | 6.5 |
| 32122 | 500 | 14.5 | 2500 | 4.0 | 2575 | 6.5 |
| 32123 | 500 | 14.5 | 2500 | 4.0 | 2600 | 6.5 |
| 32124 | 500 | 14.5 | 2500 | 4.0 | 2625 | 6.5 |
| 30638 | 1250 | 14.2 | 6250 | 4.4 | 496 | 5.4 |
| 30639 | 1250 | 14.2 | 6250 | 4.4 | 558 | 5.4 |
| 32037 | 500 | 13.1 | 2500 | 4.2 | 450 | 4.7 |
| 30694 | 1200 | 13.6 | 6000 | 4.6 | 480 | 4.5 |
| 30695 | 1200 | 13.6 | 6000 | 4.6 | 540 | 4.5 |
| 30696 | 1200 | 13.6 | 6000 | 4.6 | 600 | 4.5 |
| 30753 | 1150 | 13.1 | 5750 | 4.8 | 456 | 3.5 |
| 30754 | 1150 | 13.1 | 5750 | 4.8 | 513 | 3.5 |
| 30755 | 1150 | 13.1 | 5750 | 4.8 | 570 | 3.5 |
| 30756 | 1150 | 13.1 | 5750 | 4.8 | 627 | 3.5 |
| 30817 | 1100 | 12.5 | 5500 | 5.0 | 495 | 2.5 |
| 30818 | 1100 | 12.5 | 5500 | 5.0 | 550 | 2.5 |
| 30819 | 1100 | 12.5 | 5500 | 5.0 | 605 | 2.5 |
| 30820 | 1100 | 12.5 | 5500 | 5.0 | 660 | 2.5 |
| 30821 | 1100 | 12.5 | 5500 | 5.0 | 715 | 2.5 |
Overall score = score_ratio - 2 * repetitive DNA content
