Automated design of genomic Southern blot probes
Mike DR Croning, David G Fricker, Noboru H Komiyama and Seth GN Grant
Design Case - ProbeDesign83 - back to list
Summary
./analyse_probe_search_results : Wed Nov 9 09:33:30 2005
Connected to host: host, as user: user Database : southern_blot_designs Fetched probe design: ProbeDesign83 assembly : NCBIM34 bias : 3prime chromosome : 1 id : 74 strand : 1 ------------- Design window length: 3501bp All jobs for probe design are successful - total: 56 Fetched conf : Exonerate mouse_NCBIM34 (id: 1) Genome file(s) at: /blastdb/Mouse/NCBIM34/softmasked_dusted Fetched 1114 putative probes to analyse Selection criteria to rank probes: Minimum score ratio : 10 (i.e. self-hit must score at least 10 times higher than the next best hit) Maximum percent repetitive bases: 5 Favour probes to the 3prime end of design window ------------------------ Putative probes without hits : 0 Putative probes with single_self_hit : 1114/1114 Putative probes that are unique : 0/1114 No putative probes pass with the above criteria Now trying: score-ratio 2, and percent repetitive 40 Putative probes exceeding selection criteria : 27/1114 (both unique and not unique) Putative probes with successfully-picked primers: 26/27 Identified 1 'redundant' probes exceeding selection criteria (but wholly contained by other probes, and higher repetitive DNA content)
Image
Unique (yellow), passed (green) and failed putative probes in the design window
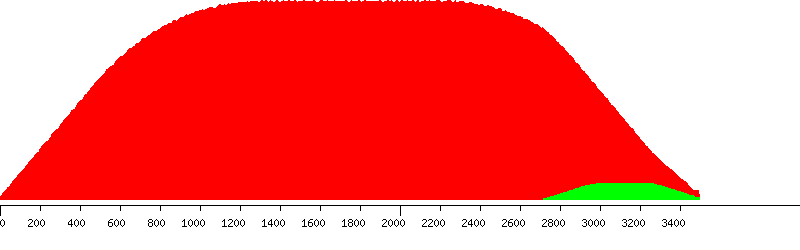
Unique putative probes
No probes
Putative probes
| Putative probe ID | Length | Score ratio | Actual | %rep DNA | Distance | Overall |
|---|---|---|---|---|---|---|
| 70282 | 500 | 15.2 | 2500 | 24.0 | 175 | -32.8 |
| 70283 | 500 | 14.9 | 2500 | 27.0 | 200 | -39.1 |
| 70281 | 500 | 11.6 | 2500 | 26.6 | 150 | -41.6 |
| 70280 | 500 | 8.0 | 2500 | 26.6 | 125 | -45.2 |
| 70279 | 500 | 6.5 | 2500 | 26.6 | 100 | -46.7 |
| 70278 | 500 | 5.2 | 2500 | 26.6 | 75 | -48.0 |
| 70171 | 550 | 13.5 | 2750 | 30.9 | 162 | -48.3 |
| 70277 | 500 | 4.9 | 2500 | 27.0 | 50 | -49.1 |
| 70170 | 550 | 10.1 | 2750 | 30.9 | 135 | -51.7 |
| 70284 | 500 | 9.5 | 2500 | 32.0 | 225 | -54.5 |
| 70172 | 550 | 8.3 | 2750 | 31.6 | 189 | -55.0 |
| 70276 | 500 | 4.7 | 2500 | 30.8 | 25 | -56.9 |
| 70169 | 550 | 7.6 | 2750 | 33.3 | 108 | -58.9 |
| 70166 | 550 | 5.2 | 2750 | 32.2 | 27 | -59.2 |
| 70165 | 550 | 5.2 | 2750 | 32.5 | 0 | -59.9 |
| 70168 | 550 | 6.0 | 2750 | 33.3 | 81 | -60.5 |
| 70167 | 550 | 5.4 | 2750 | 33.3 | 54 | -61.1 |
| 70072 | 600 | 9.2 | 3000 | 36.7 | 120 | -64.1 |
| 70073 | 600 | 8.0 | 3000 | 36.7 | 150 | -65.3 |
| 70071 | 600 | 7.0 | 3000 | 36.7 | 90 | -66.4 |
| 70173 | 550 | 6.4 | 2750 | 36.5 | 216 | -66.7 |
| 70275 | 500 | 4.7 | 2500 | 35.8 | 0 | -66.9 |
| 70074 | 600 | 6.0 | 3000 | 36.7 | 180 | -67.3 |
| 70068 | 600 | 5.7 | 3000 | 37.8 | 0 | -70.0 |
| 70069 | 600 | 5.7 | 3000 | 38.2 | 30 | -70.7 |
| 70070 | 600 | 6.1 | 3000 | 38.8 | 60 | -71.6 |
Overall score = score_ratio - 2 * repetitive DNA content
Redundant probes
| Putative probe ID | Length | Score ratio | Actual | %rep DNA | Distance | Overall |
|---|---|---|---|---|---|---|
| 70285 | 500 | 6.7 | 2500 | 37.0 | 250 | -67.3 |
Overall score = score_ratio - 2 * repetitive DNA content
